Single cell is here!
Today, we're incredibly excited to announce the launch of Pluto's end-to-end analysis experience for single cell RNA-seq. Single cell RNA sequencing (single cell RNA-seq or scRNA-seq) is a technique that has steadily gained massive popularity over the past few years. Unlike traditional bulk RNA-seq, single cell RNA-seq analyzes the transcriptional landscape within individual cells, uncovering intricate details about gene expression and cell-to-cell variability.
By considering the transcriptome at a single cell resolution, scientists can unravel complex biological processes, uncover rare cell types, delineate developmental trajectories, and decode heterogeneity within tissues, offering new insights into the fundamental workings of living organisms. At Pluto, we're thrilled to make it easier and faster than ever before for scientists to gain insight from complex scRNA-seq data.
~ Caitlin Winkler, PhD | Lead Solutions Scientist at Pluto
Analyzing scRNA-seq data with Pluto
Designed from the ground up to be biologist-friendly and intuitive, Pluto's single cell RNA-seq analysis experience empowers any scientist on your team to upload raw data, run QC and alignment pipelines, perform flexible preprocessing steps, annotate clusters of cell types, and perform downstream differential analysis.
Data upload (FASTQs) -> QC and alignment pipeline
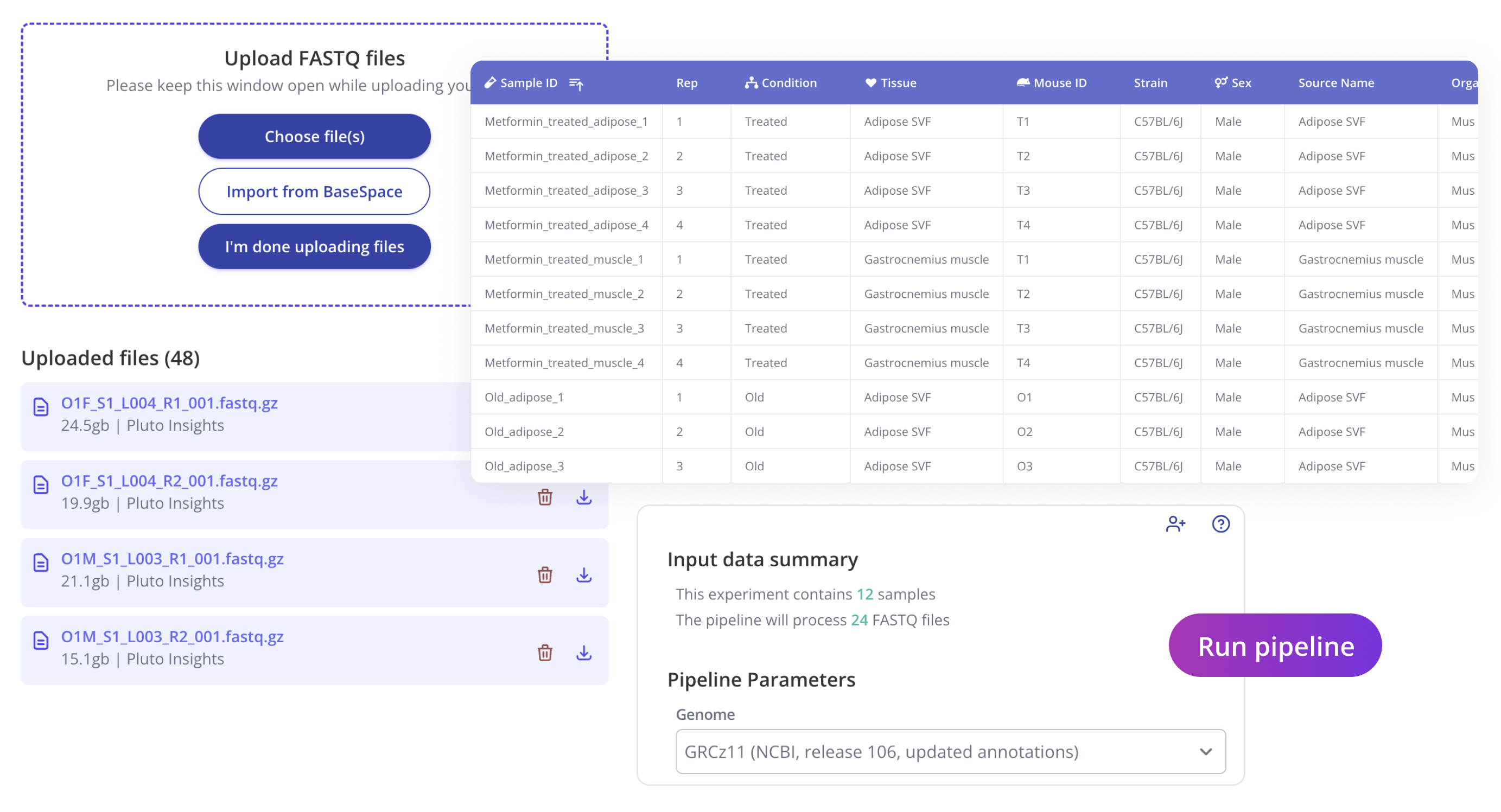
You'll begin your journey similar to other NGS assays in Pluto, by uploading FASTQ files and sample annotations. Pluto's sample annotation table can contain any biological metadata relevant to your experiment and will be used to enable flexible downstream analysis later on. Once your data has been uploaded, select the desired genome build for alignment and run the pipeline.
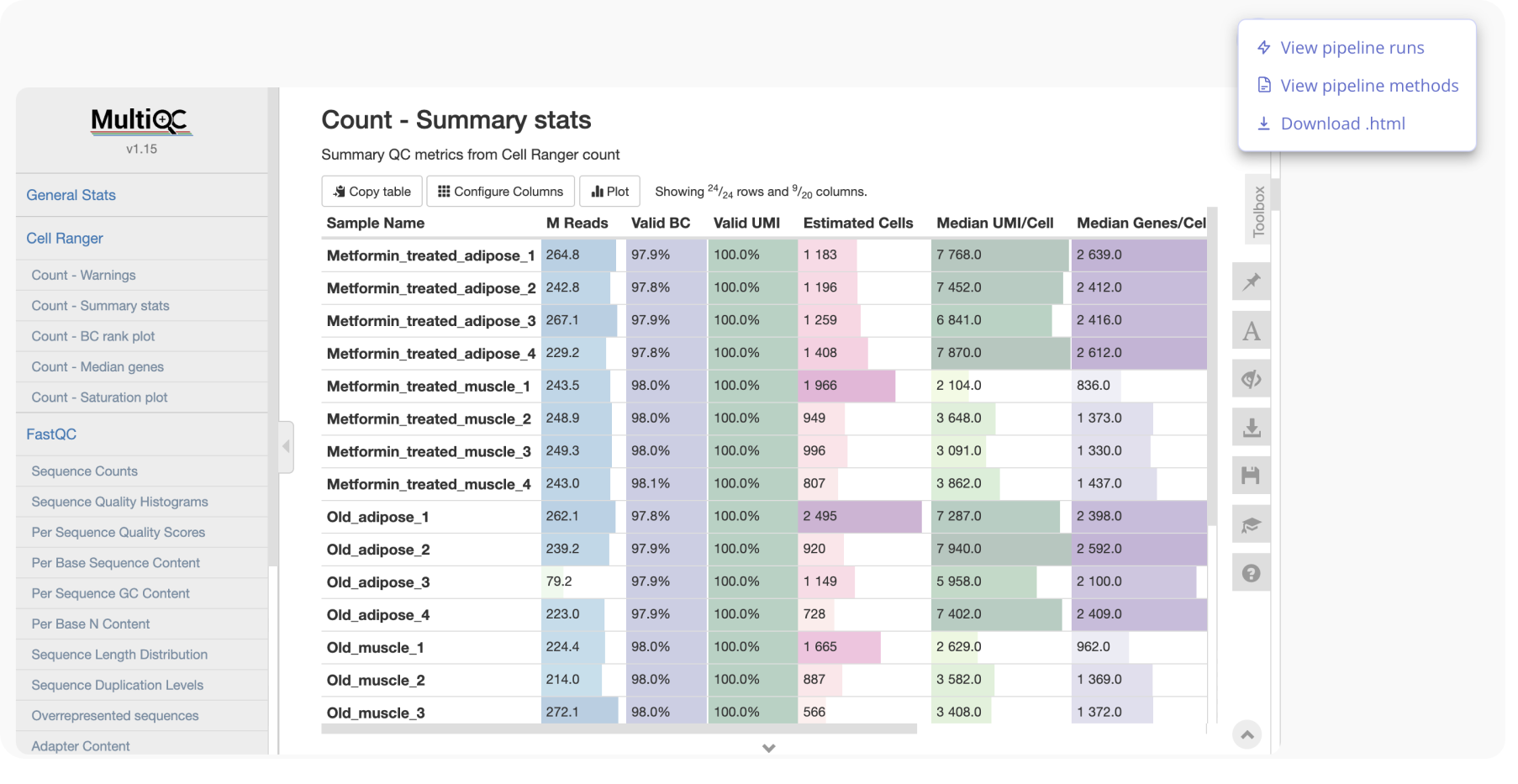
You and any collaborators shared on the experiment will get an email notification when the pipeline has completed! At that time, you can review the MultiQC report and pipeline methods.
Pre-processing: filtering, normalization, integration, clustering
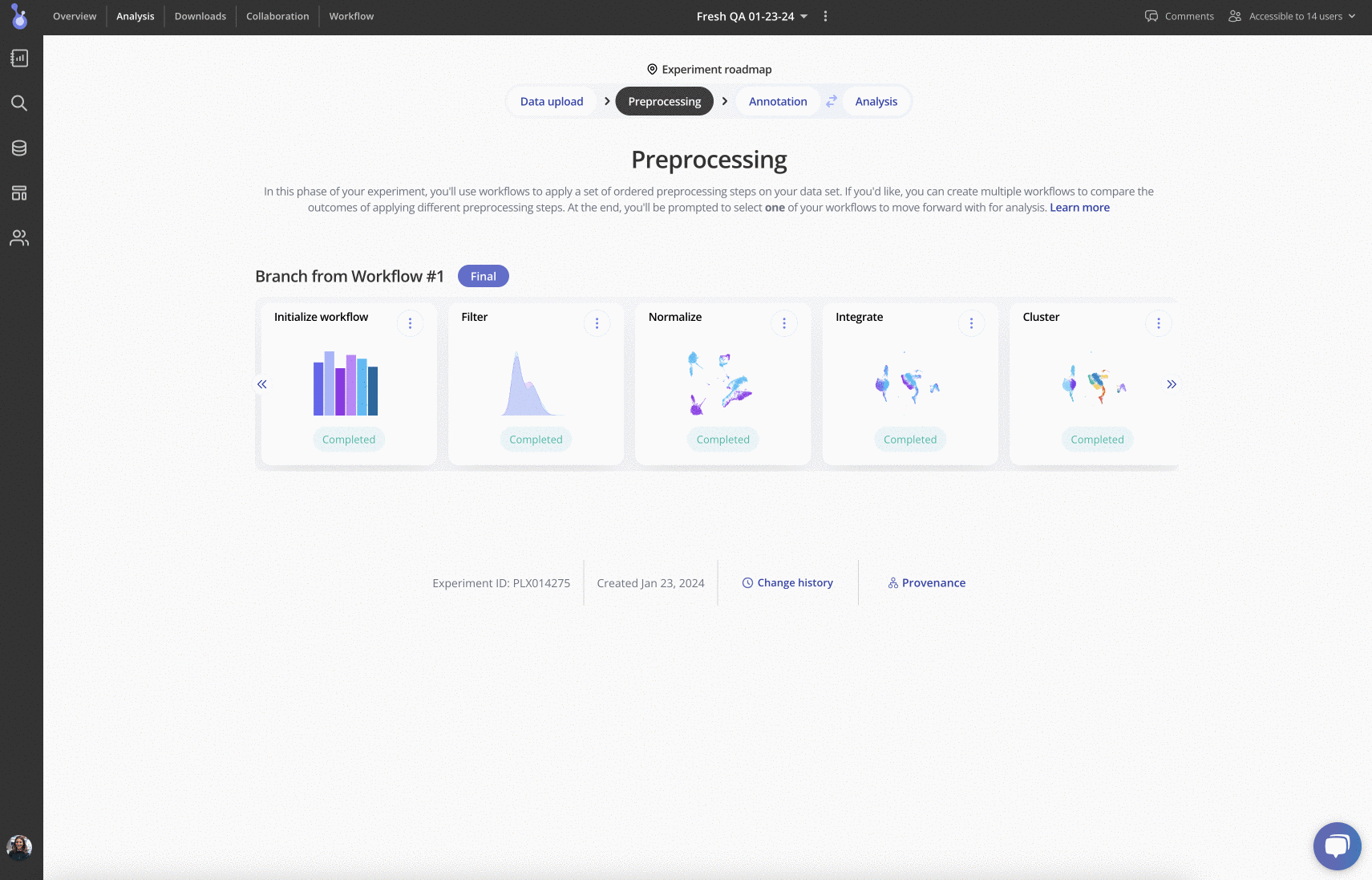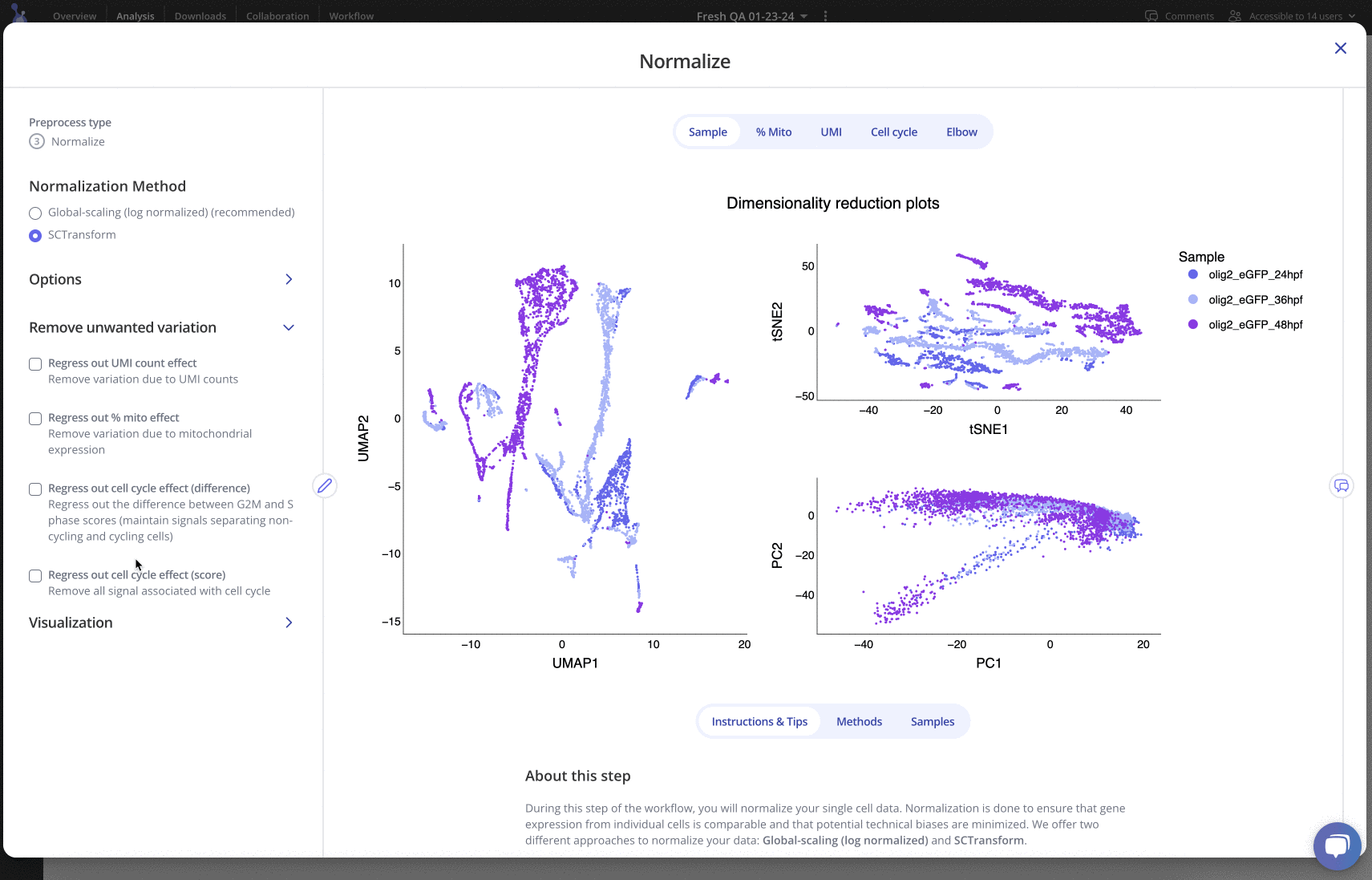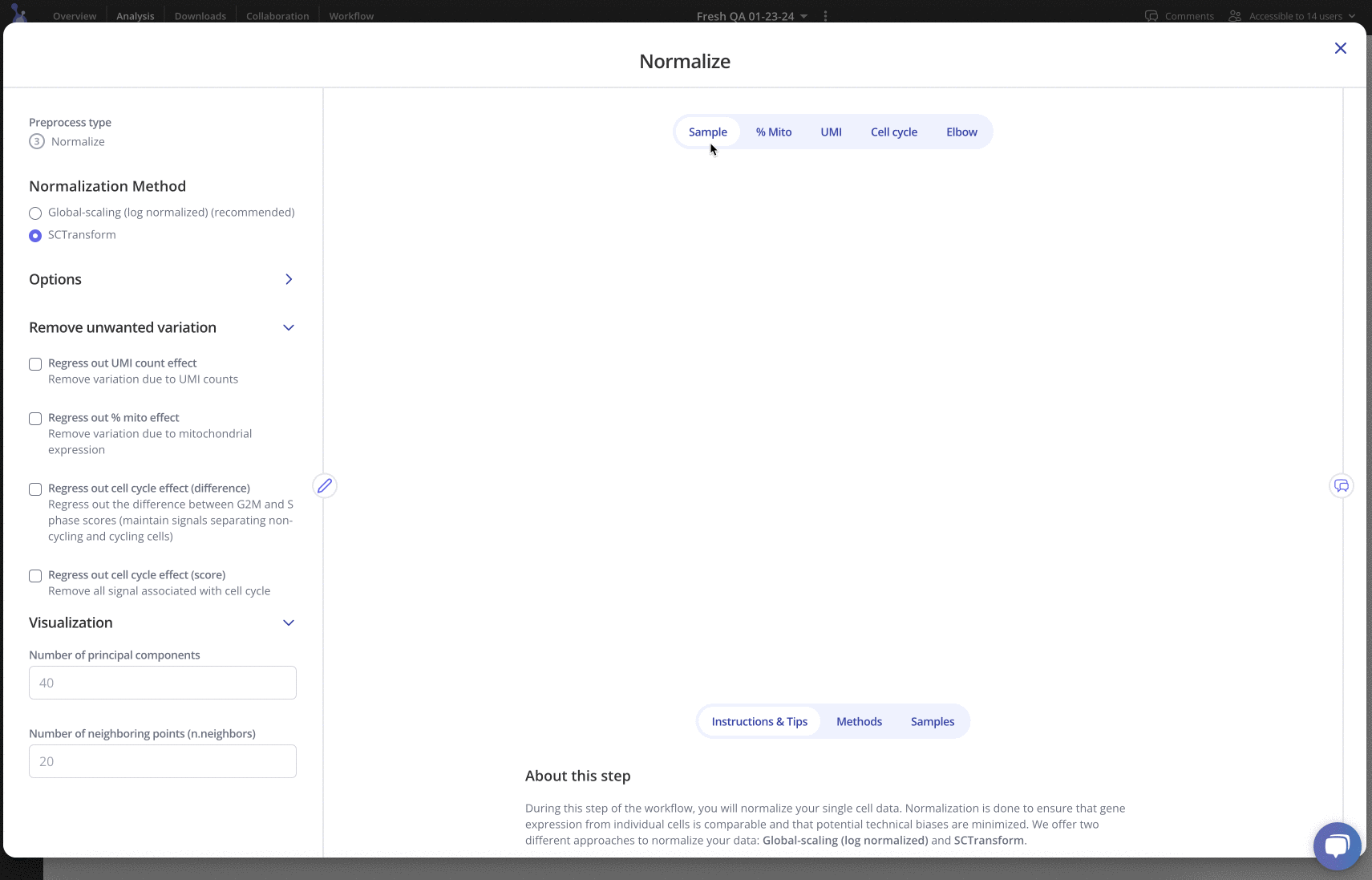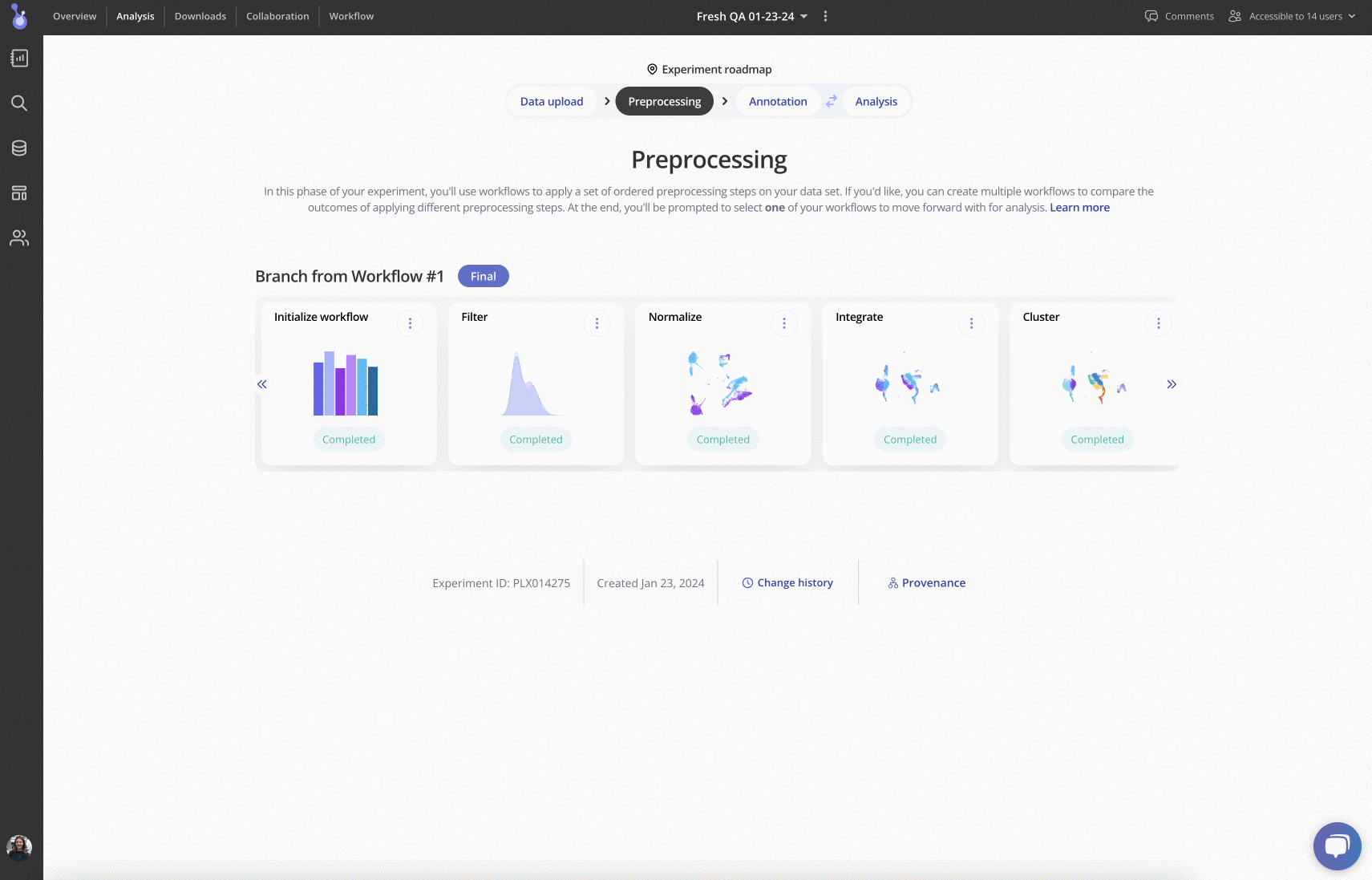
Up next is pre-processing: in this phase, you can try out different workflows for performing filtering, normalization, integration, clustering and more. Each pre-processing step has a variety of parameters that can be edited, plus plots and tables to be explored:
For more information, check out the detailed vignettes on various pre-processing stages in Pluto:
Cell type annotation
In the pre-processing phase, you can select one or multiple resolutions to carry forward into analysis. On the annotation page, you can manage different sets of labels or annotations for the clusters in each of those resolutions.
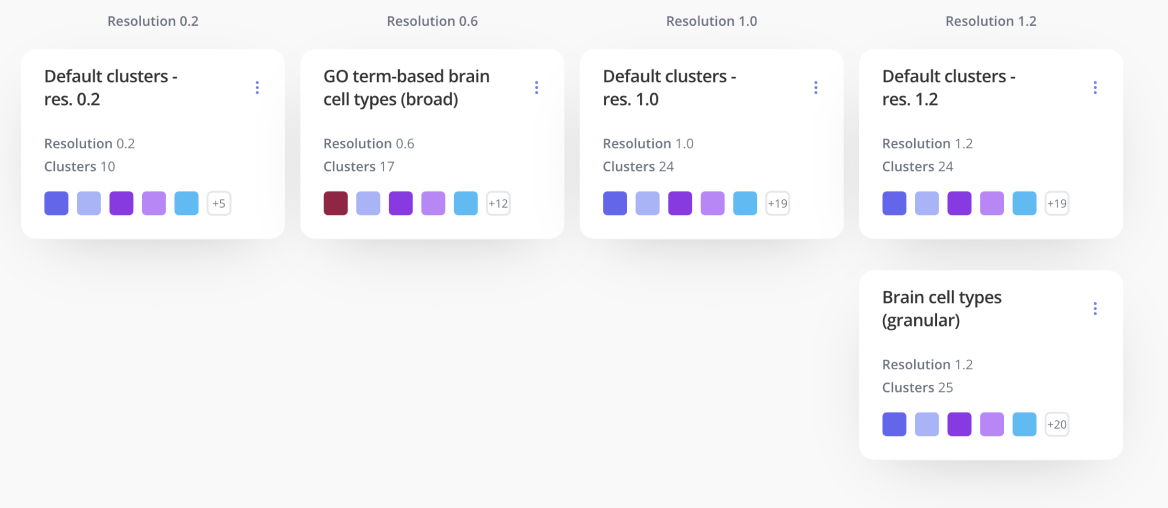
For any of your label sets, you can investigate top marker genes, annotate clusters, include rationale and methods, and customize colors in an easy-to-use interface. Learn more
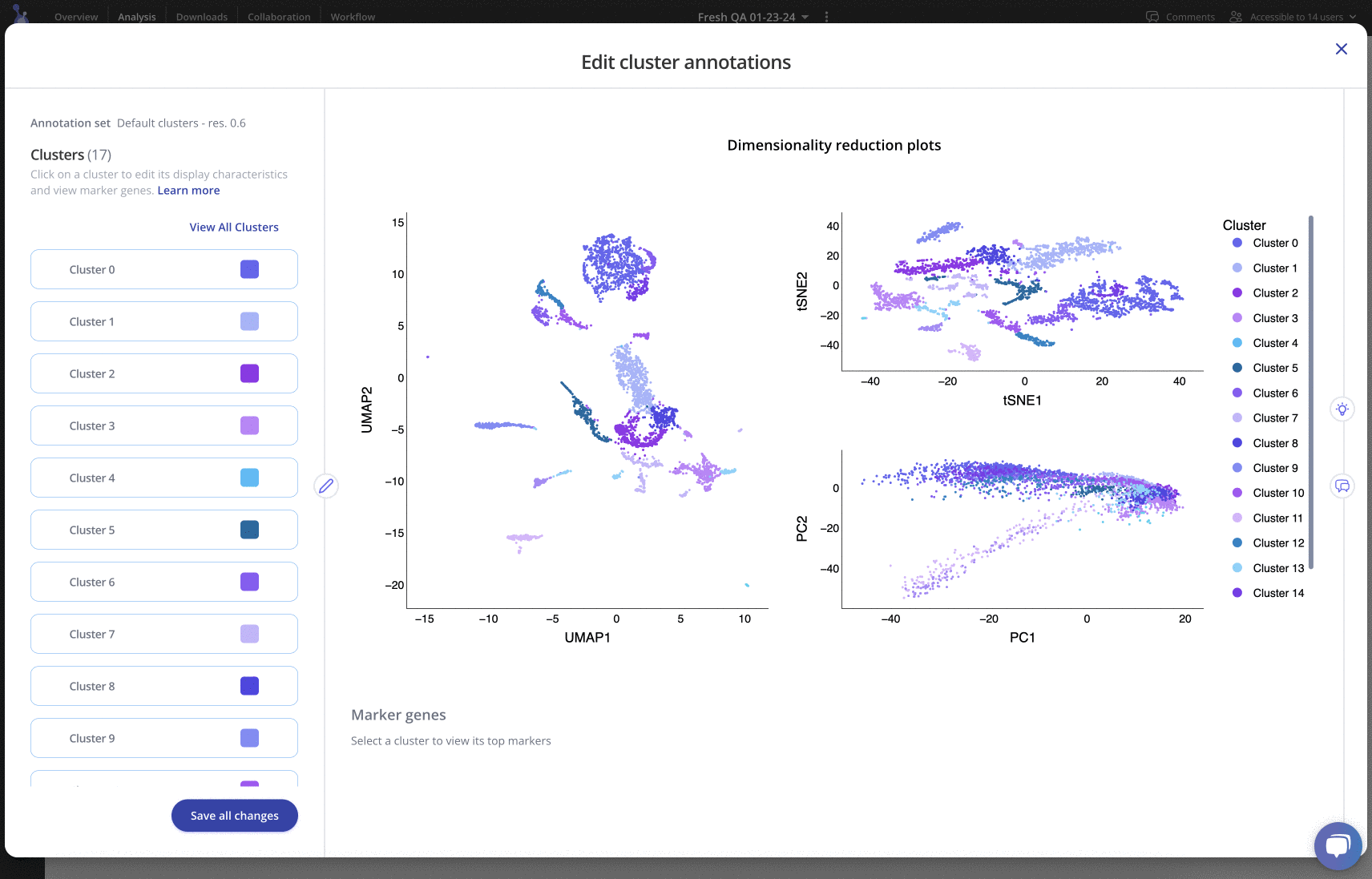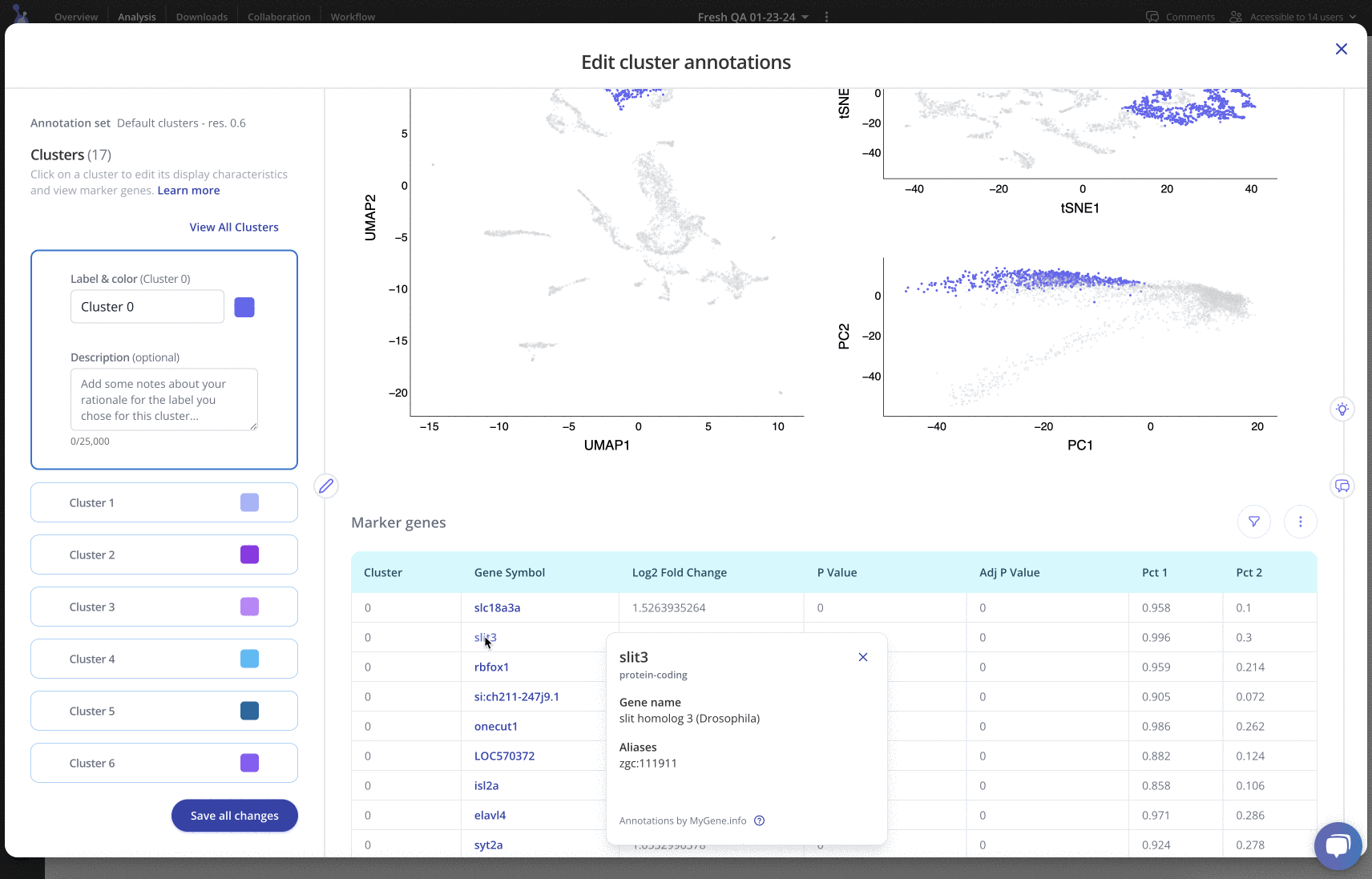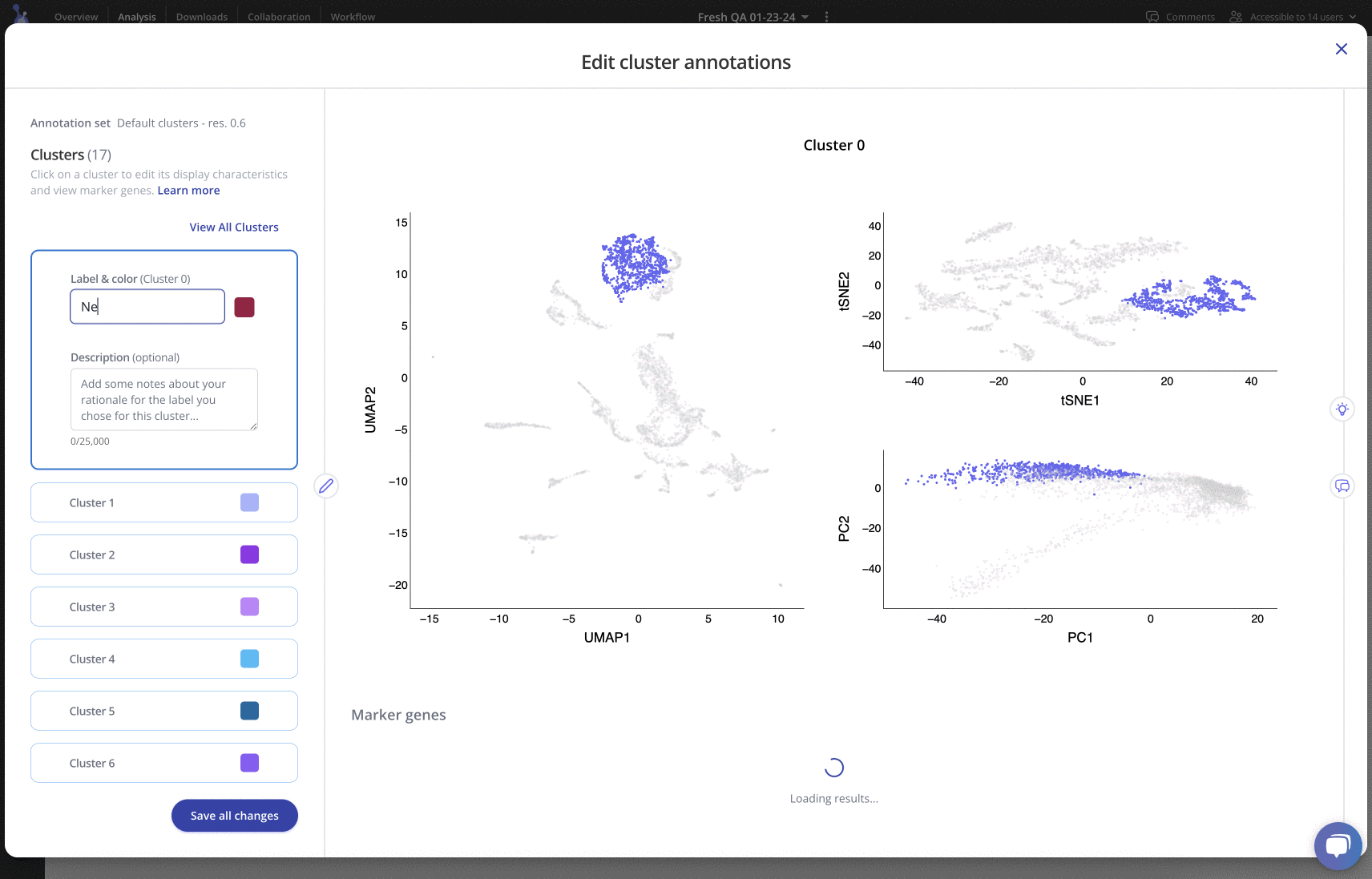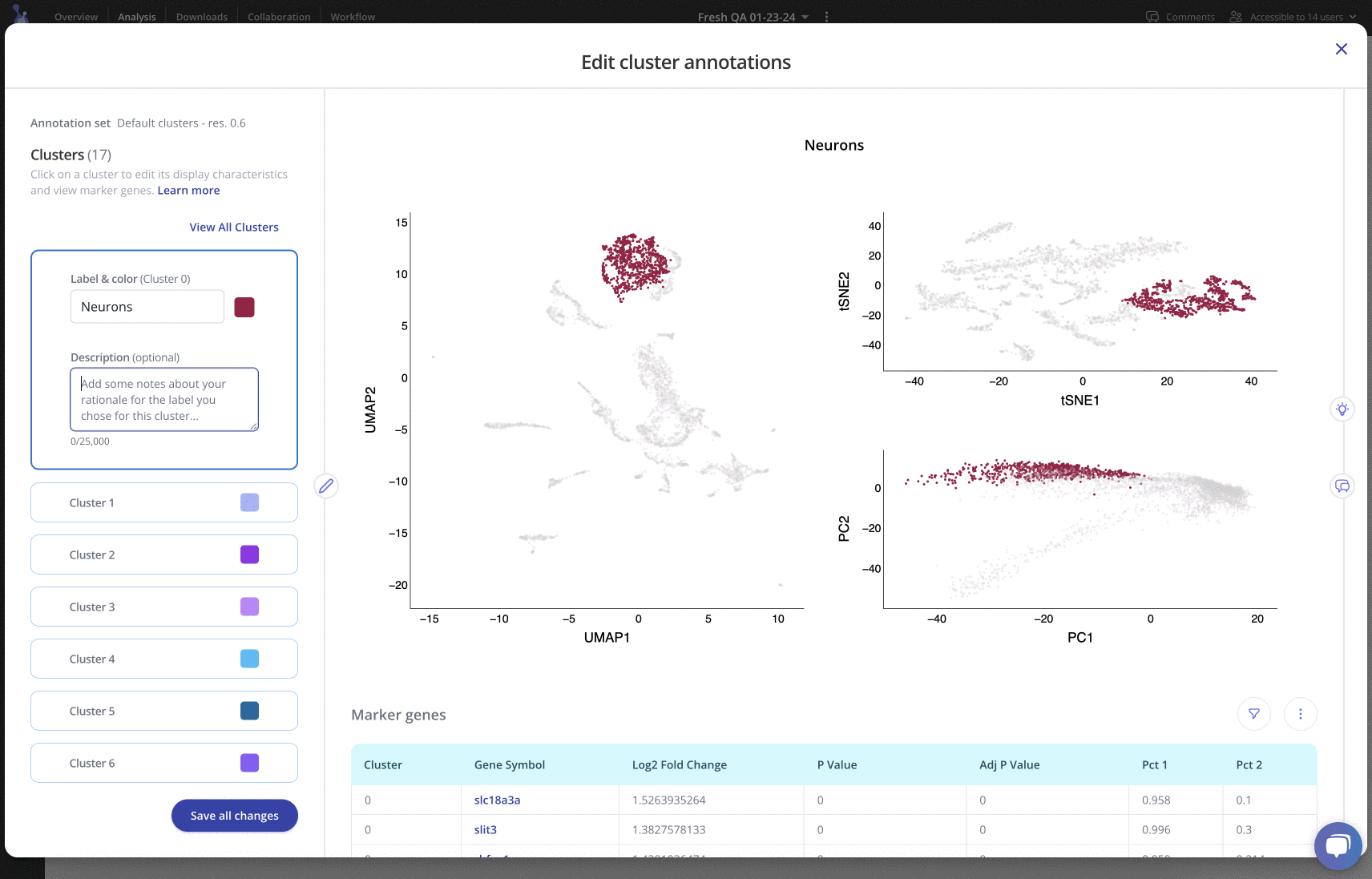
Downstream analysis
Finally, to interrogate biological questions of interest, perform downstream analysis. With Pluto's flexible interface, you can create comparisons by cluster, biological condition (e.g. timepoint, treatment), investigate doublets/singlets, compare cells at different phases of the cell cycle and more!
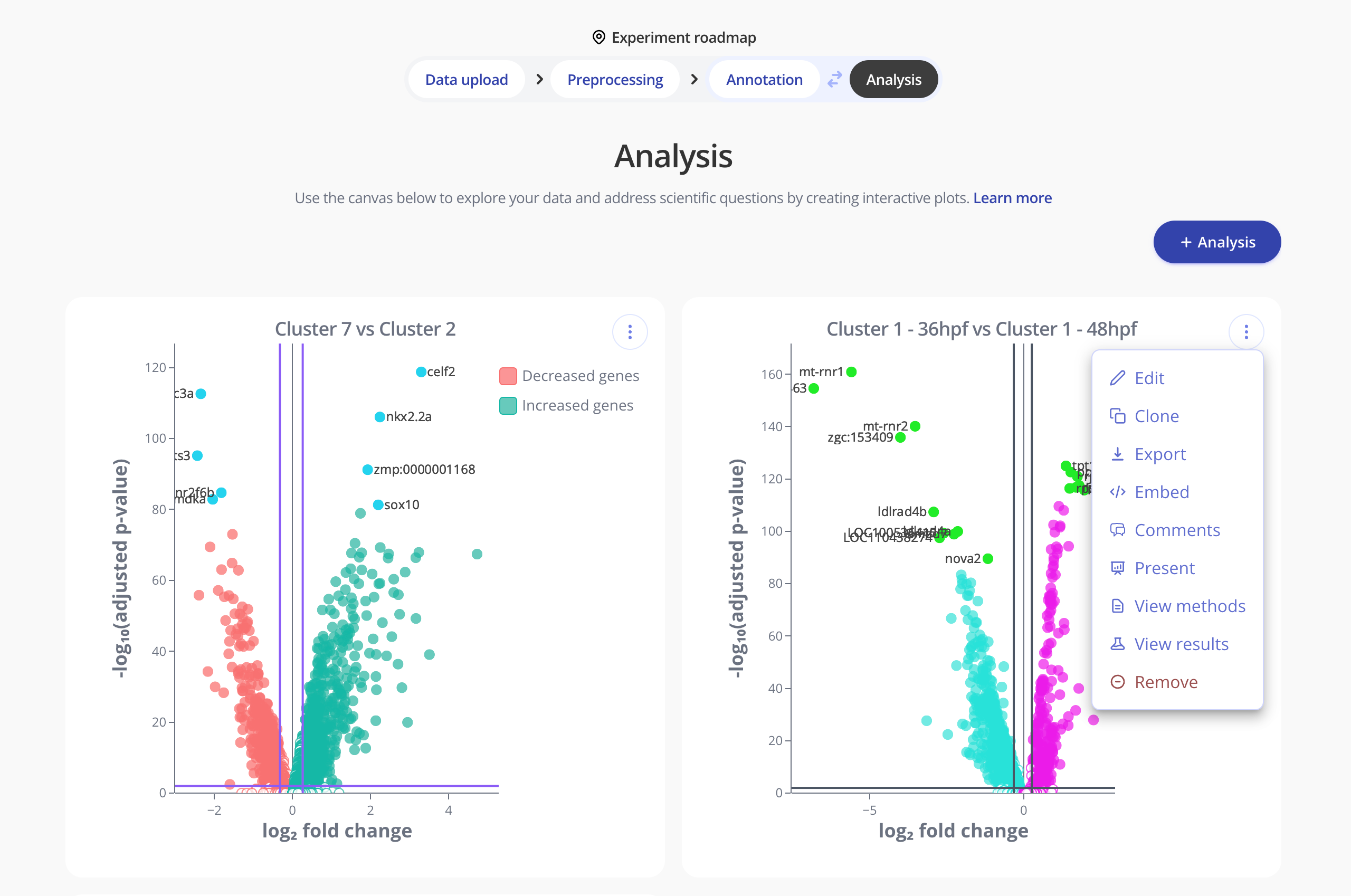
Ready to try it for yourself?
Join the high-velocity scientific teams already using Pluto to accelerate their discoveries from scRNA-seq and other assays. Contact us for personalized demo of the entire platform.

