We’re proud to highlight a new paper citing Pluto published today in Cells by Dr. Holger Russ’ lab at the Barbara Davis Center for Diabetes, University of Colorado School of Medicine.
Type 1 diabetes (T1D) is an autoimmune disease where the patient's own immune system destroys their insulin-producing beta cells in error. Currently, patients must rely on endogenous insulin injections as treatment. Dr. Holger Russ and his team study the underlying mechanisms that lead to the development of T1D, with the goal of helping pioneer better treatment options.
About this study
In their latest publication, published in the journal Cells, the Russ Lab investigates the role of the PTPN2 gene in T1D. PTPN2 is a known, high-risk T1D-associated gene expressed in both immune- and pancreatic beta cells, but how genes affect the development of autoimmune diabetes is largely unknown.
Previously, Dr. Russ was among the first to demonstrate the generation of functional beta cells from human pluripotent stem cells under cell culture conditions. Now, the team is taking advantage of recent breakthroughs in genome editing technology to establish different inducible CRISPR/Cas9 systems to facilitate rapid and precise gene modification of pluripotent stem cells and their recently differentiated derivatives.
Dr. Russ summarized the impact of this study:
In this study we used stem cell technology and CRISPR/Cas9 genome engineering technology to inactivate the T1D risk gene PTPN2 in stem cell derived beta cells. We found that such cells upregulate expression of genes important for immune surveillance. Using innovative co-culture of autoreactive T cells with PTPN2-/- beta cells revealed increased immune activation compared to controls. This provides novel insights how T1D risk genes could impact immune beta cell interactions and expands our understanding of potential diseases mechanisms.
Creating and customizing the figures in Pluto
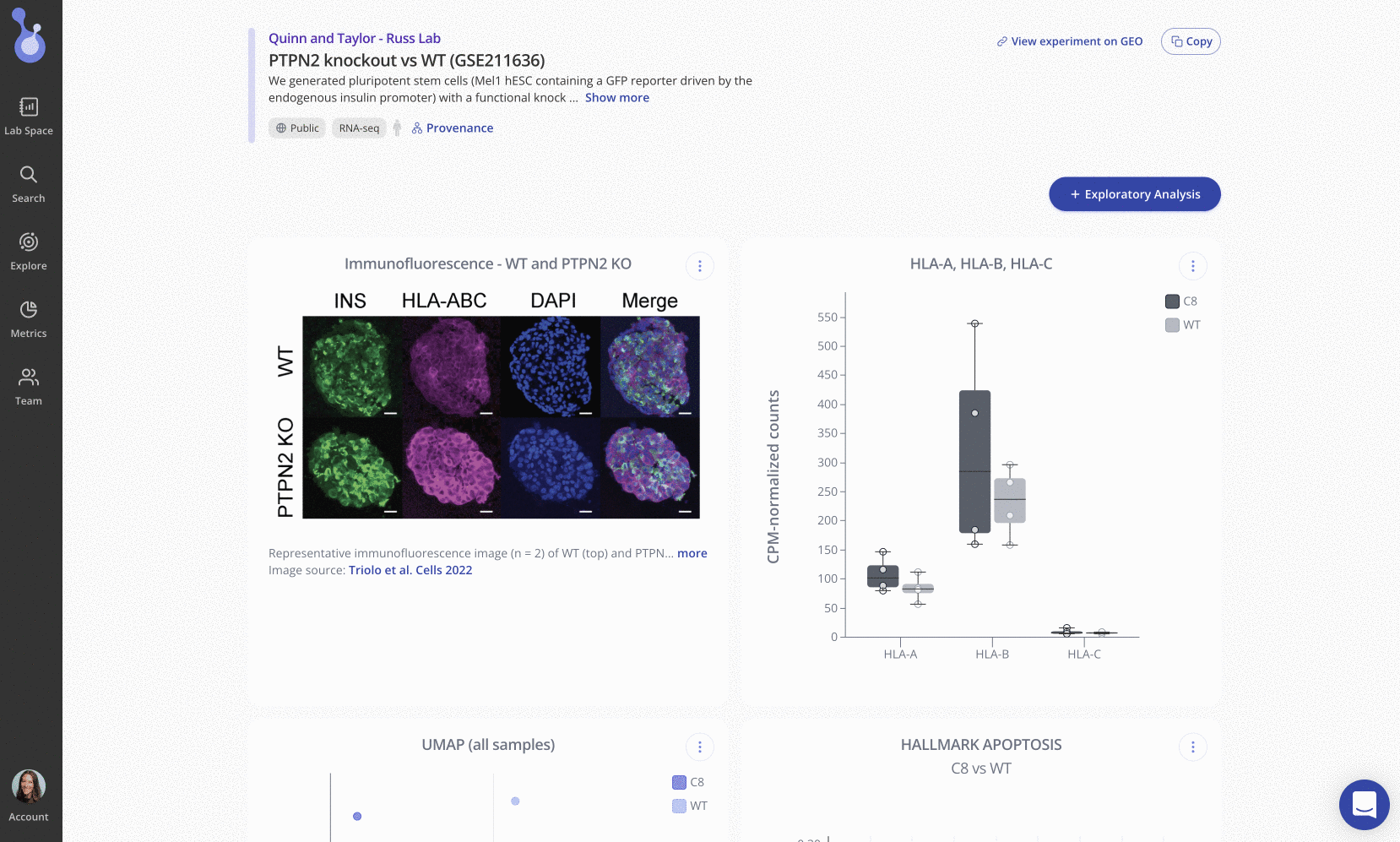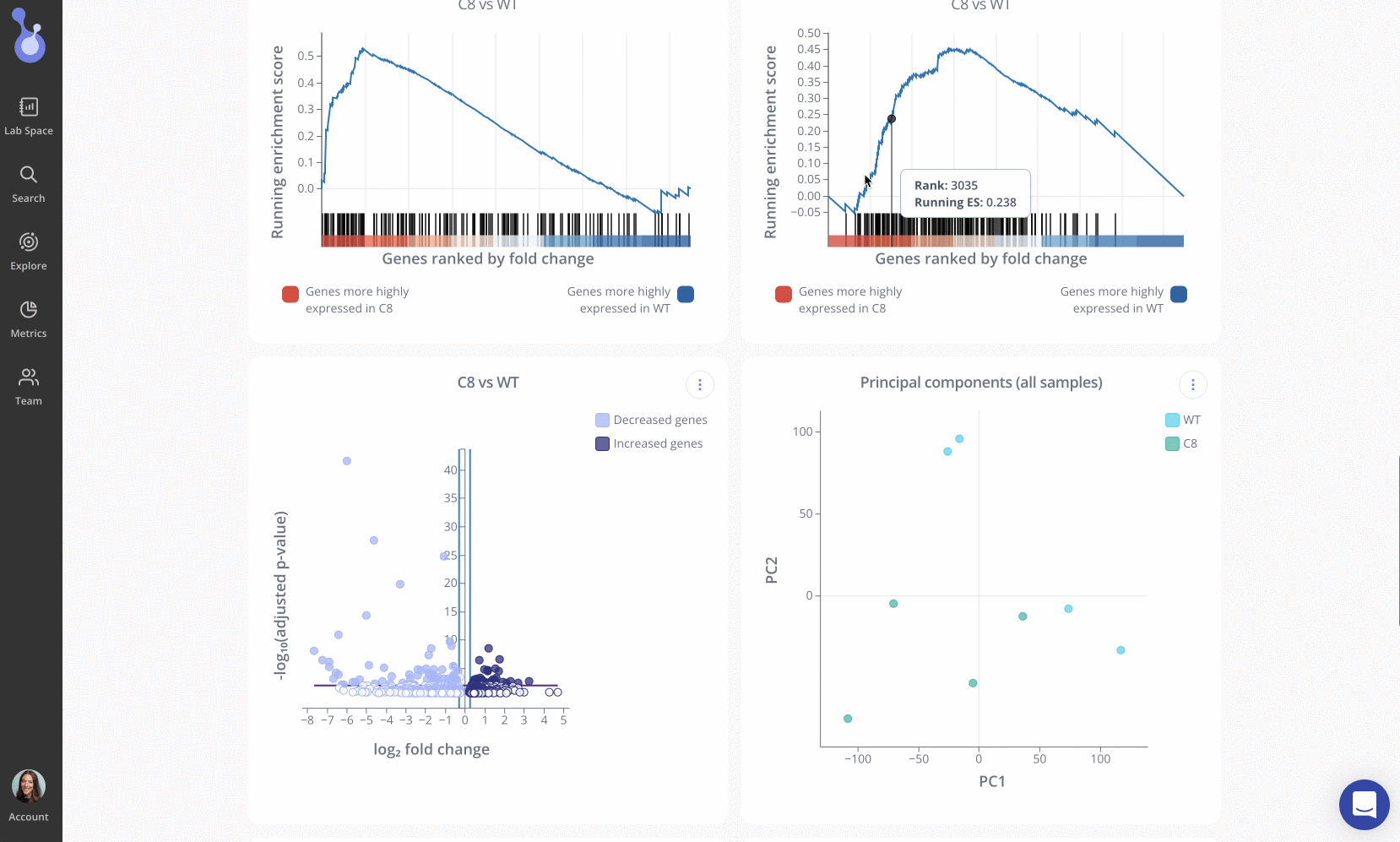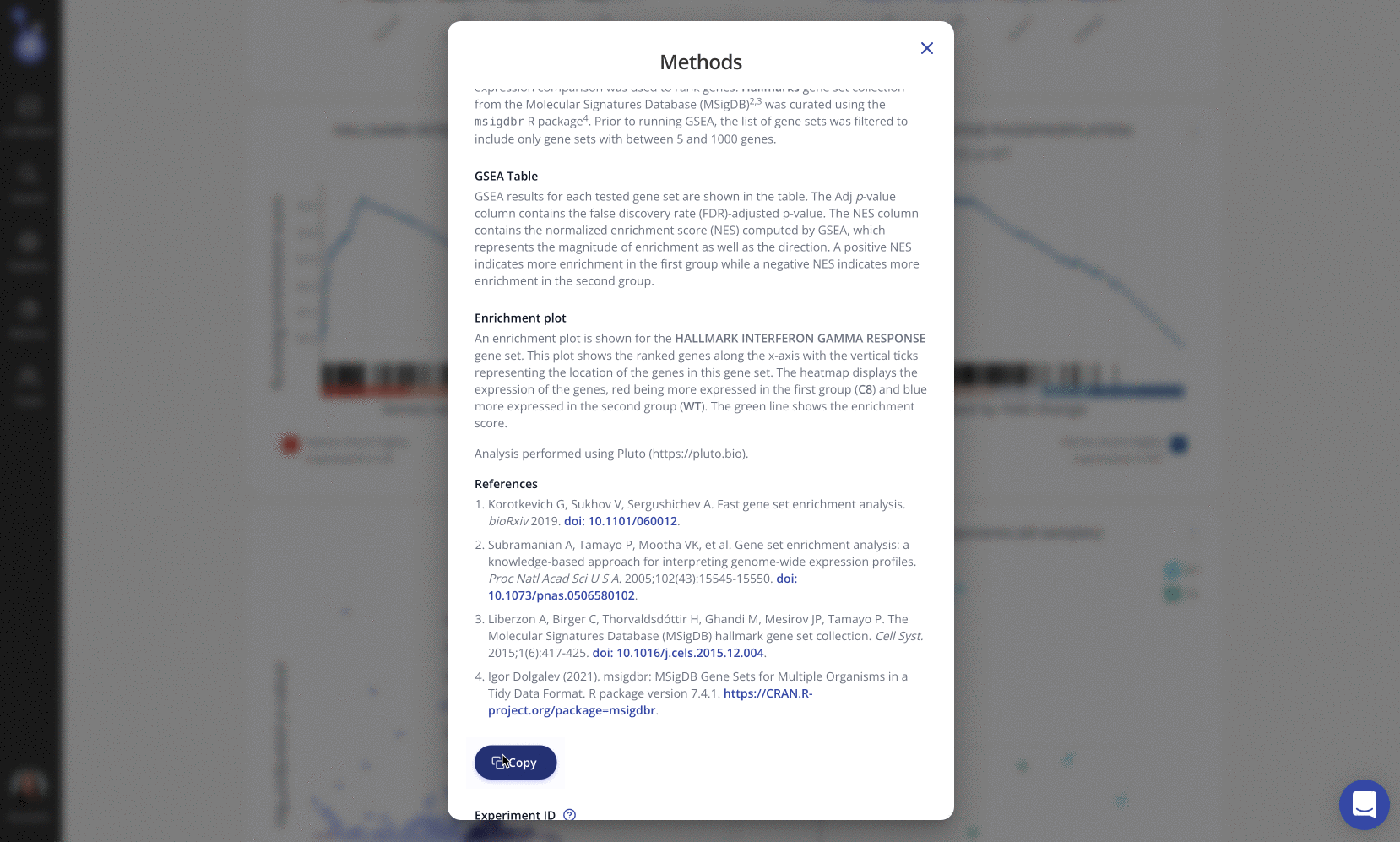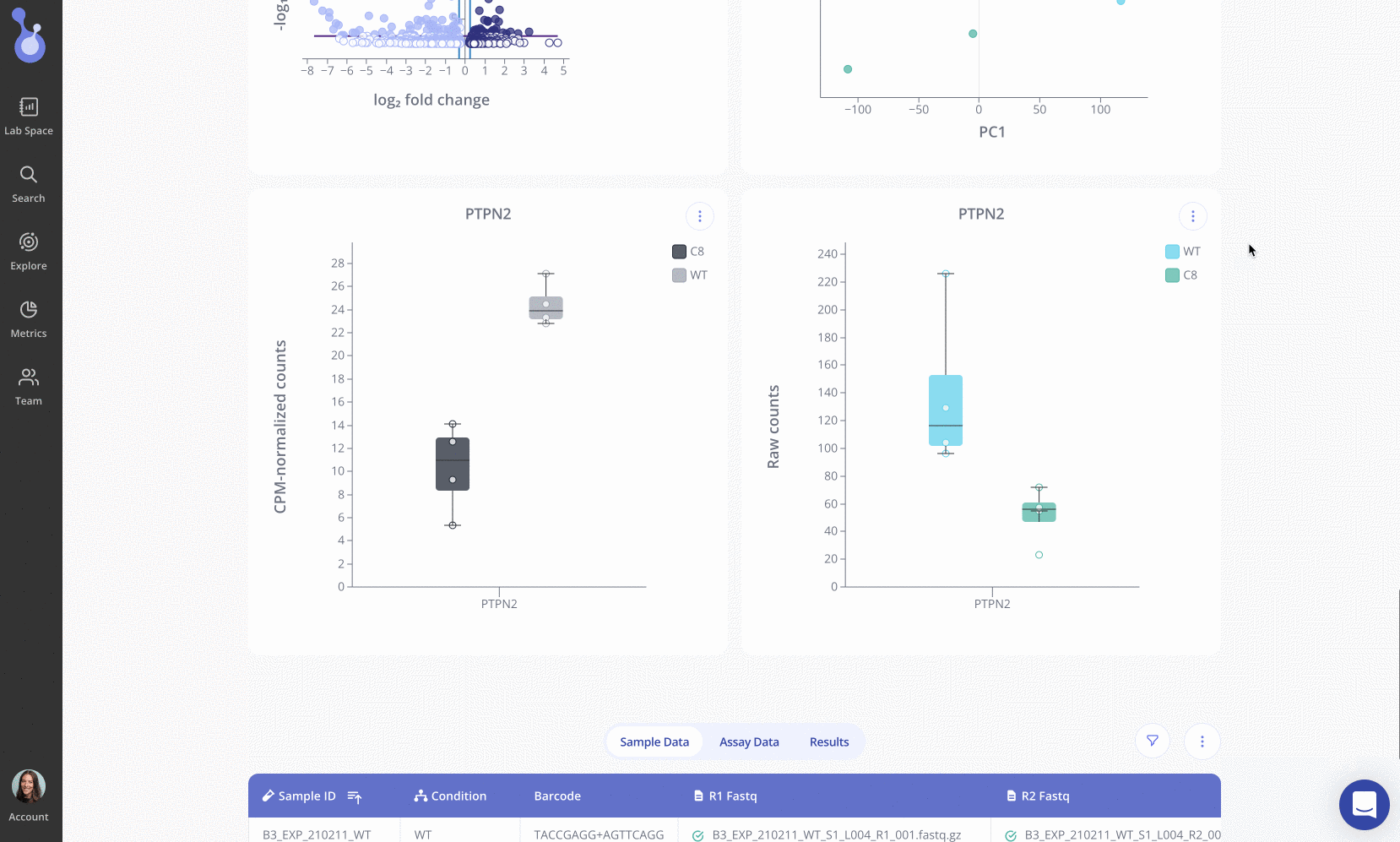
After using CRISPR/Cas9 to generate a functional knockout of PTPN2 in human pluripotent stem cells (hPSC) and differentiating stem-cell-derived beta-like cells (sBC), the team measured genome-wide gene expression in the sBCs using bulk RNA sequencing.
Using Pluto’s intuitive, web-based interface, Russ Lab members Taylor Triolo and Quinn Matuschek uploaded their bulk RNA-seq data to Pluto in FASTQ format for processing via Pluto’s automated RNA-seq pipeline. After pre-processing, alignment, and quality control, Taylor and Quinn performed differential expression analysis to identify 308 differentially expressed genes in the PTPN2 knockout line compared to WT control samples.
Note: the plot above shows live data from Pluto for this experiment. Click the button in the upper right corner to view the methods. Learn how to create your own public plot share links with Pluto here.
Next, the team performed gene set enrichment analysis (GSEA), which revealed alterations in the Hallmark IFNγ response, oxidative phosphorylation, and IL6 JAK/STAT3 signaling pathways.
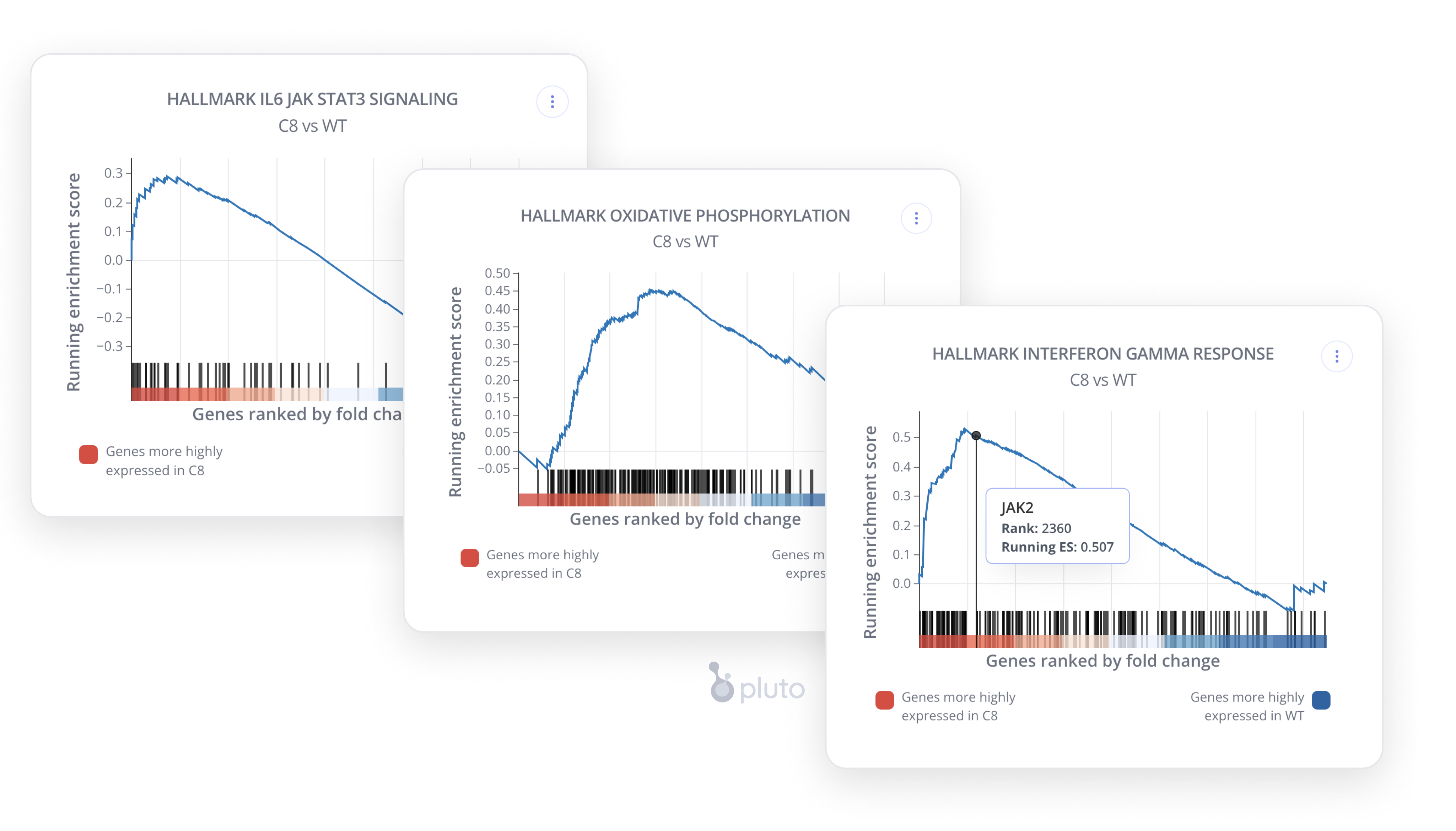
We loved the Russ Lab's choice of a simple and consistent color palette for their figures, which was easily customized by editing the Pluto plot settings. Pluto also automatically generates the dynamic methods section so the Russ Lab could paste it in their manuscript without additional effort.
The end result? An impactful publication and a fully interactive page with rich data and results!
The Pluto team congratulates the Russ Lab on these novel and clinically relevant findings! Be sure to check out the publication in Cells.
References
Triolo, T. et al. Stem-Cell-Derived β-Like Cells with a Functional PTPN2 Knockout Display Increased Immunogenicity. Cells 11(23), 3845. 2022. https://doi.org/10.3390/cells11233845
