Introduction
RNA sequencing (RNA-seq) has revolutionized the field of genomics by providing a powerful tool for quantifying gene expression levels. This technology allows researchers to investigate differential gene expression under different conditions, providing valuable insights into various biological processes and diseases. However, to draw accurate and meaningful conclusions from RNA-seq data, it is essential to consider covariates, which are additional, potentially “hidden” variables that may impact gene expression in your experiment. In this blog, we’ll discuss the definition, role, and importance of including covariates in differential expression analysis when analyzing RNA-seq data.
What are covariates?
Covariates, also known as confounding variables, are factors other than the main independent variable(s) of interest that can affect the outcome being studied. In the context of RNA-seq data analysis, covariates can be various biological, technical, or environmental factors that may influence gene expression levels. Common covariates include age, sex, batch effects, disease status, and treatment conditions1.
The role of covariates in differential expression analysis
Differential expression analysis is a fundamental task in RNA-seq data analysis, where researchers aim to identify genes that are differentially expressed between two or more experimental conditions. The presence of covariates introduces complexity into this analysis because they can confound the interpretation of differential expression results. Some key roles that covariates play in differential expression analysis include:
-
Controlling for confounding effects: Covariates can mask or exaggerate the true differences in gene expression between experimental groups. By including covariates in the analysis, researchers can statistically control for their effects, ensuring that observed differences are not solely attributable to these confounding variables.
-
Reducing batch effects: RNA-seq experiments often involve multiple batches or sequencing runs. Batch effects can introduce unwanted variability in gene expression data. Covariates can help adjust for these technical differences, making it easier to identify biologically relevant differences between conditions1.
-
Enhancing statistical power: By accounting for covariates, the statistical power of the analysis is improved. This means that researchers are more likely to detect true differential expression events while minimizing the risk of false positives.
Why include covariates?
The inclusion of covariates is critical for the accuracy and robustness of differential expression analysis in RNA-seq data. Here are some reasons why covariates should not be overlooked:
-
Biological relevance: Covariates often represent biologically relevant factors that can provide deeper insights into the biological processes under investigation. When we're asking a scientific question like "How does gene expression change in patients with an infection compared to healthy uninfected controls," it’s important to be aware that biological factors such as patient age or sex could impact their gene expression, and thus may be reflected in your differential analysis results if not accounted for. Ignoring covariates may lead to erroneous conclusions and hinder the understanding of complex biological systems.
-
Data quality control: Covariates can help identify and correct for potential sources of bias or technical variation in the data, ensuring that the results reflect true biological differences rather than experimental artifacts.
-
Reproducibility: Research findings must be reproducible to be considered valid. Including covariates in your analysis pipeline enhances the reproducibility of RNA-seq experiments by accounting for sources of variability that may not be immediately apparent.
-
Regulatory standards: In some research fields, such as clinical genomics, regulatory agencies like the FDA may have recommendations for the inclusion of covariates to ensure statistical robustness and higher confidence2,3.
Including covariates in differential expression analysis
Most standard tools and packages for performing differential expression analysis include a method for accounting for covariates. When writing your own R scripts with the DESeq2 package, for example, you can include covariates in the design matrix to be passed to the DESeq2 function4.
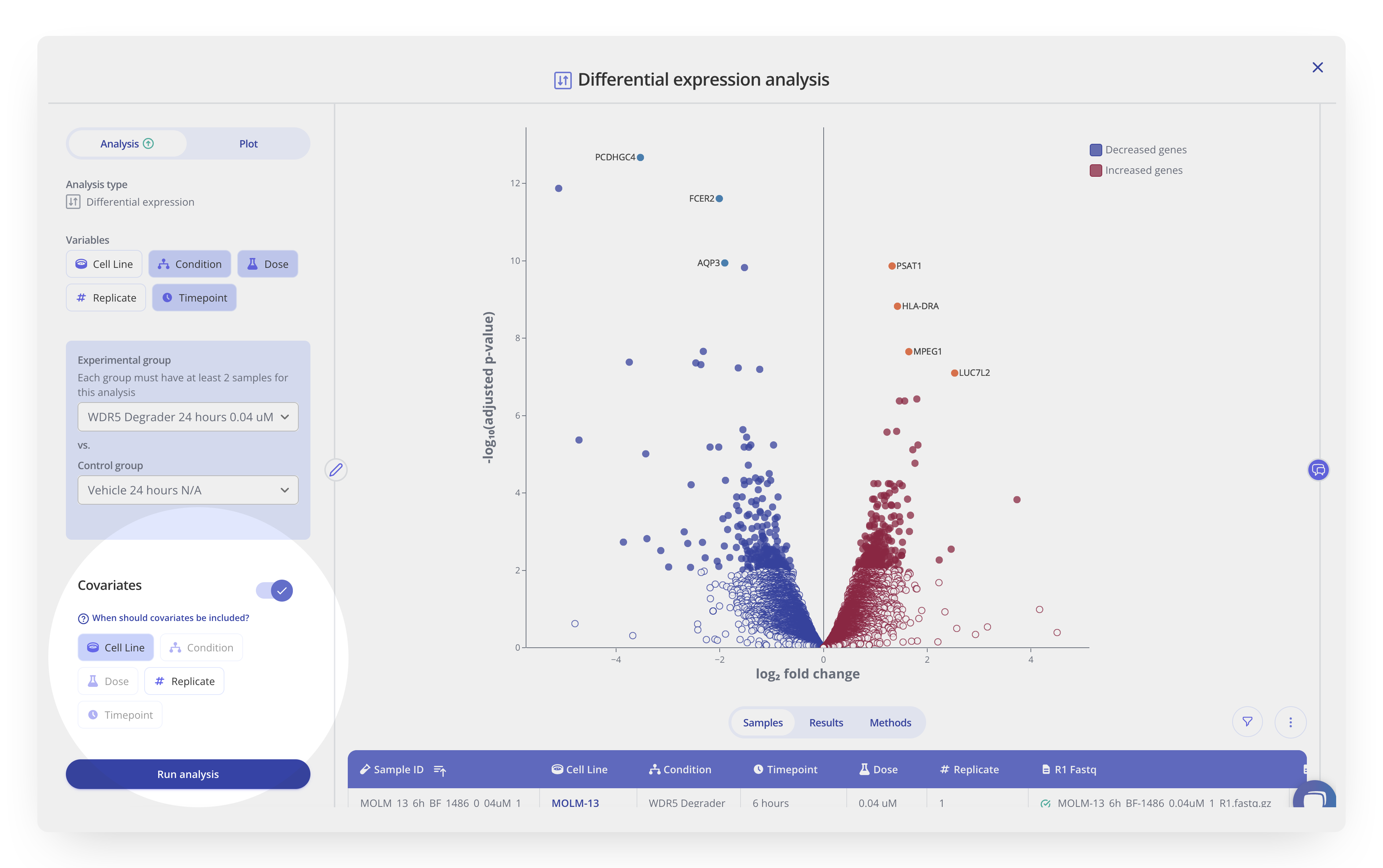
In Pluto, you can include covariates in your analysis without needing to write any additional code. Toggle on the Covariates option and select one or more potentially confounding factors to include as covariates in your analysis. Quickly test different combinations of covariates by cloning the analysis and editing which variables are included. Importantly, any modifications made to the covariates in an analysis will remain automatically up-to-date in the dynamic methods section available in Pluto. Learn more about performing differential analysis with covariates in Pluto.
Conclusion
In conclusion, covariates are essential components of RNA-seq data analysis, particularly in differential expression analysis. They play a crucial role in controlling for confounding effects, reducing technical variability, and enhancing the overall quality and reliability of the results. Researchers should carefully consider and include relevant covariates in their analysis pipelines to ensure that their findings accurately reflect the biology they aim to study. Consider providing covariates in your own scripts or apply them via Pluto’s intuitive analysis platform in order to uncover meaningful insights into gene expression patterns and advance our understanding of complex biological processes and diseases.
References
- Leek JT, Storey JD. Capturing Heterogeneity in Gene Expression Studies by Surrogate Variable Analysis. PLoS Genet 3(9): e161. 2007. https://doi.org/10.1371/journal.pgen.0030161.
- FDA issues final guidance on adjusting covariates in randomized clinical trials. May 31, 2023. https://www.fda.gov/drugs/drug-safety-and-availability/fda-issues-final-guidance-adjusting-covariates-randomized-clinical-trials.
- Guidance Document: Adjusting for Covariates in Randomized Clinical Trials for Drugs and Biological Products. May 25, 2023. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/adjusting-covariates-randomized-clinical-trials-drugs-and-biological-products.
- Michael I. Love, Simon Anders, and Wolfgang Huber. Analyzing RNA-seq data with DESeq2. November 14, 2023. https://bioconductor.org/packages/devel/bioc/vignettes/DESeq2/inst/doc/DESeq2.html.

