Pluto Bioinformatics
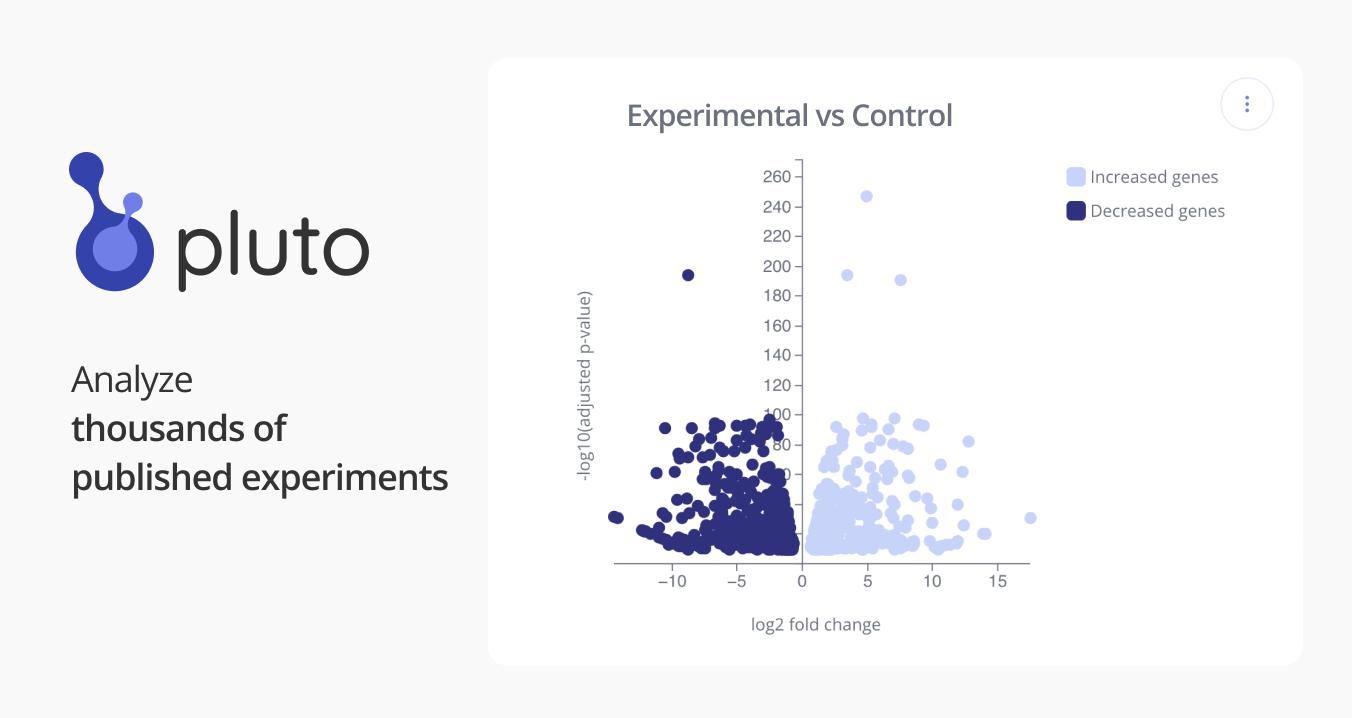
GSE129240: Complete deconvolution of cellular mixtures based on linearity of transcriptional signatures
Bulk RNA sequencing
Difference in RNA content of different cell types introduces bias to gene expression deconvolution methods. If ERCC spike-ins are introduced into samples, predicted proportions of deconvolution methods can be corrected SOURCE: Maxim Artyomov (martyomov@wustl.edu) - Artyomov lab WASHINGTON UNIVERSITY IN ST LOUIS