Pluto Bioinformatics
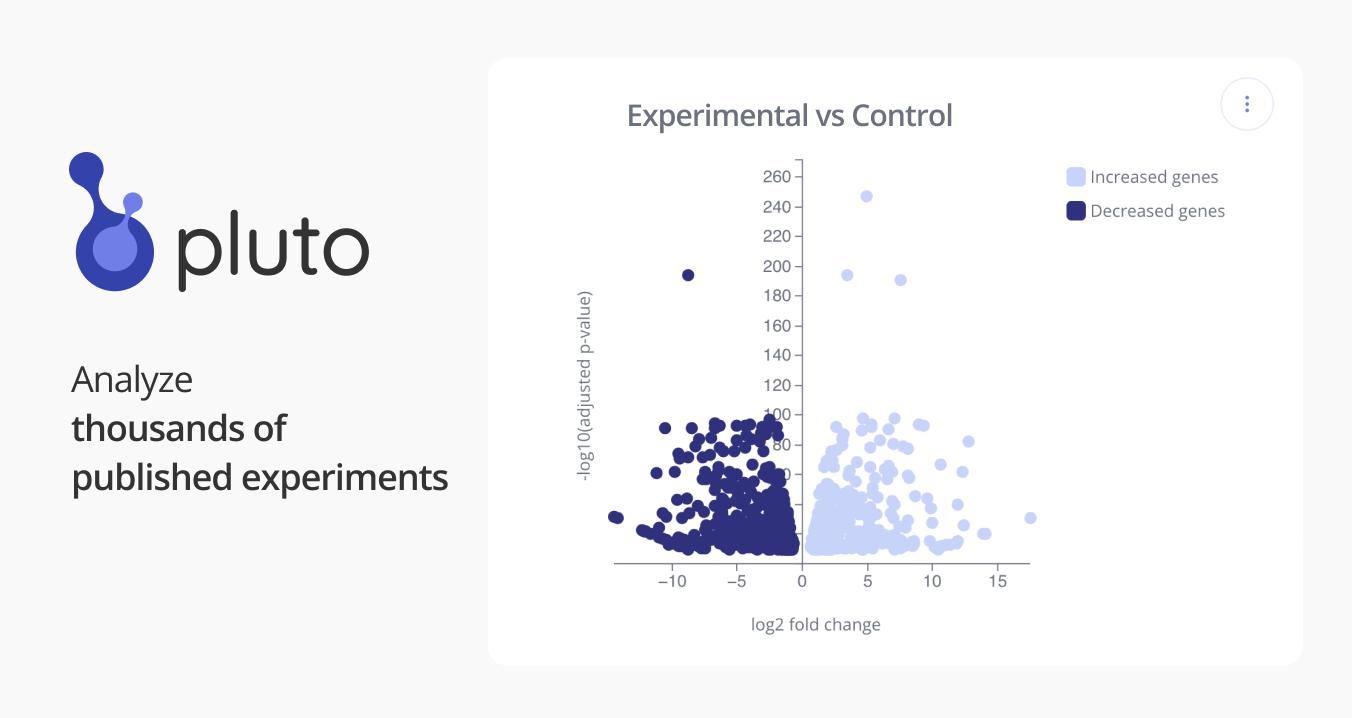
GSE90059: RNAseq analysis of AML cells upon lncRNA knockdown
Bulk RNA sequencing
It is well understood how proteins regulate cell fate, both in normal development and disease. However, a substantial fraction of the genome is transcribed in a cell type- specific manner, producing long non-coding RNAs (lncRNA) rather than protein- coding transcripts. Here we systematically characterize transcriptional dynamics (both mRNA and lncRNA) during hematopoiesis and in hematological malignancies. We present de novo assembled transcriptome models and expression values for hematopoietic lncRNAs. We found lncRNAs to be regulated during differentiation and misregulated in disease. We assessed lncRNA function via an in vivo RNAi screen in a model of acute myeloid leukemia. With this approach, we identified several lncRNAs essential for leukemia maintenance, and found that a number act by promoting leukemia stem cell signatures. Leukemia blasts show a myeloid differentiation phenotype when these lncRNAs were depleted, and our data indicates that this effect is mediated via effects on the c-MYC oncogene. SOURCE: M Joaquina Delas (mjdelasv@cshl.edu) - Cold Spring Harbor Laboratory