Pluto Bioinformatics
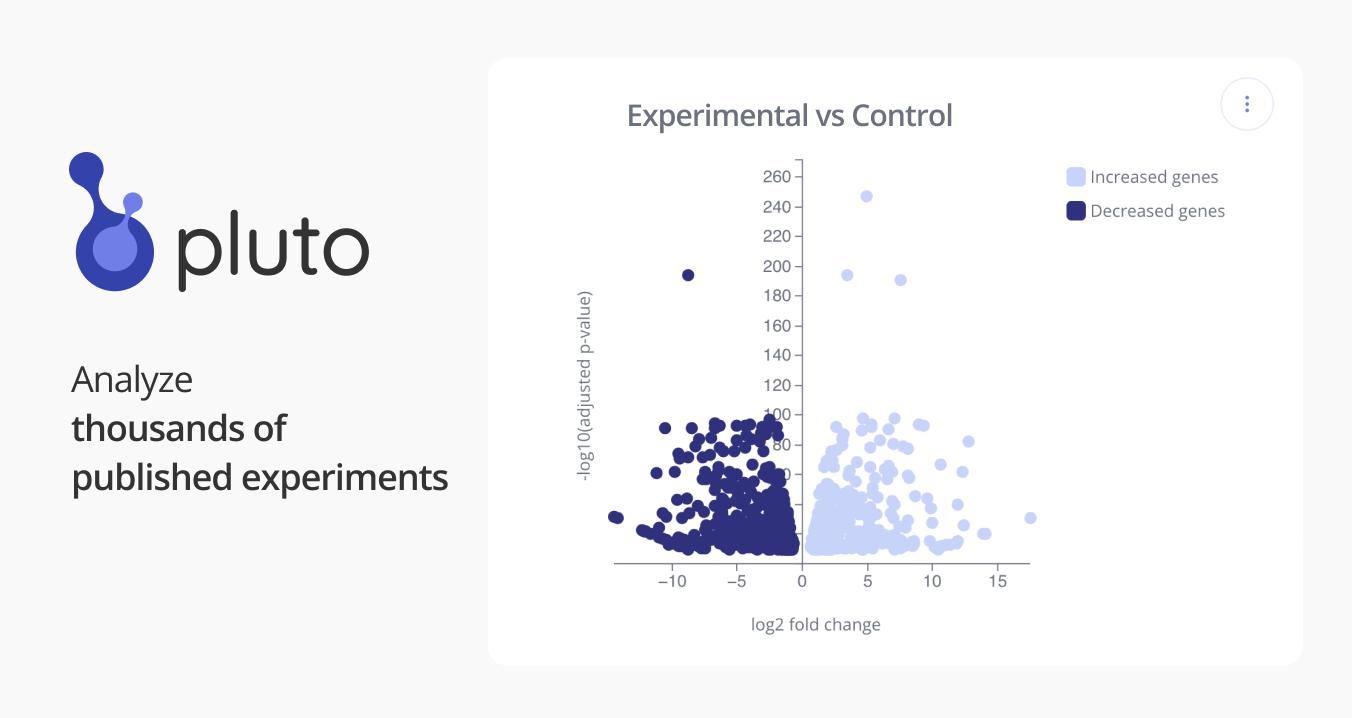
GSE107942: Lipoxins Regulate the Egr-1 Network and Reverse Diabetic Kidney Disease
Bulk RNA sequencing
The failure of spontaneous resolution underlies chronic inflammatory conditions including microvascular complications of diabetes such as diabetic kidney disease. The identification of endogenously generated molecules which promote the physiologic resolution of inflammation suggests that these bioactions may have therapeutic potential in the context of chronic inflammation. Lipoxins (LXs) are lipid mediators which promote the resolution of inflammation. Here we investigated the potential of LXA4 and a synthetic LX analogue [Benzo-LXA4] as therapeutics in a murine model of diabetic kidney disease. Diabetes was induced with streptozotocin in ApoE-/- mice and kidney disease was observed. The development of diabetes-induced albuminuria, mesangial expansion and collagen deposition was attenuated by LXs. It is noteworthy that LXs also attenuated established kidney disease in diabetic mice administered LXs 10 weeks after disease onset, with evidence of preserved kidney function. Kidney transcriptome profiling defined a diabetic signature [725 genes; FDR P0.05]. Comparison of this murine gene signature with human DKD identified shared renal pro-inflammatory/pro-fibrotic signals (TNF-, IL-1, NF-B). In diabetic mice we identified 20 and 51 transcripts regulated by LXA4 and Benzo-LXA4, respectively, and pathway analysis identified established (TGF-1, PDGF, TNF-, NF-B) and novel (Early growth response-1 - EGR-1) networks activated in diabetes and regulated by LXs. Using human renal epithelial cells, we demonstrate that LXs attenuate TNF- driven Egr-1 activation, with Egr-1 important for cellular responses to TGF-1 and TNF-. These data demonstrate for the first time that LXs can reverse established diabetic complications and support a therapeutic paradigm to promote the resolution of inflammation. SOURCE: Mark,D,Ziemann (mark.ziemann@gmail.com) - Epigenetics in Human Health and Disease Monash University