Pluto Bioinformatics
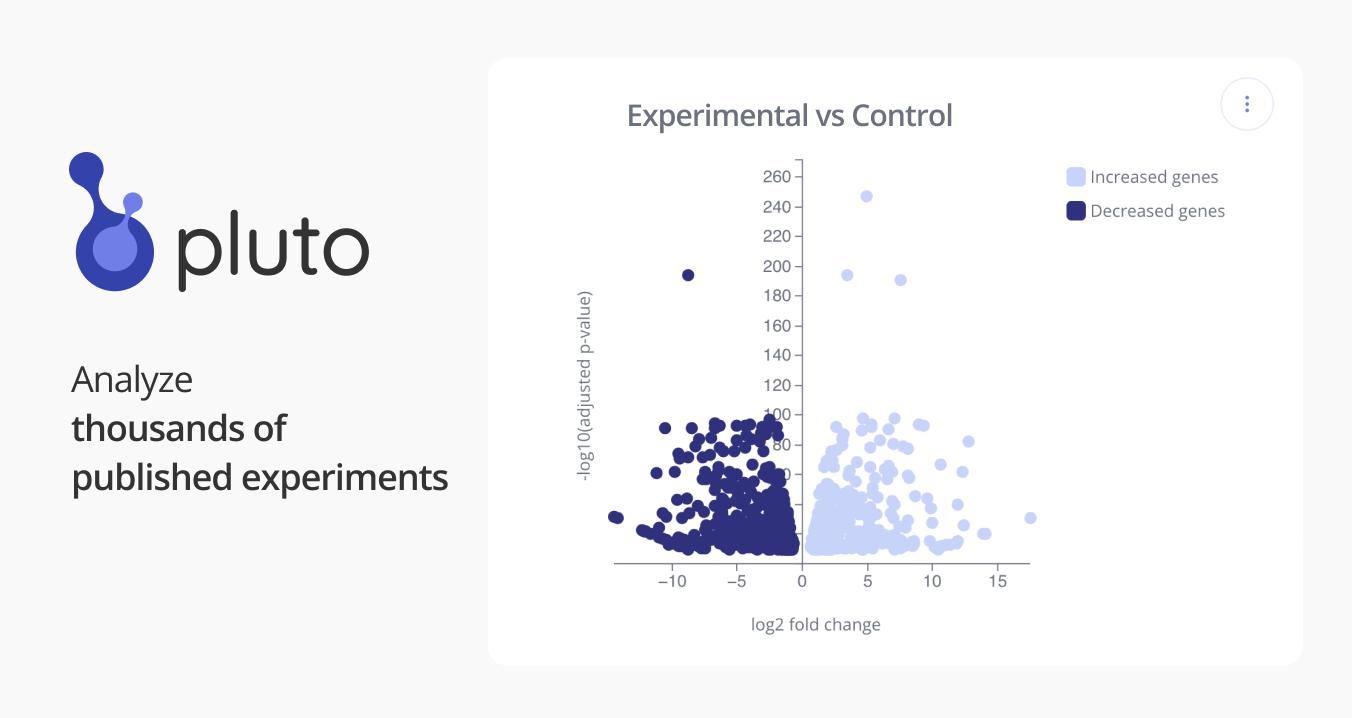
GSE110139: Defining CBX7-dependent chromatin architecture with rapid small-molecule inhibition
Bulk RNA sequencing
While the repressive roles of Polycomb complexes have been well-established with in vitro systems, it remains unclear how subtypes of these complexes function in living cells. Conditional alleles have enabled studies of Polycomb Repressive Complex 1 (PRC1), but the available timescales on which inducible deletion occurs is not sufficient to define its immediate regulatory targets. We have used rapid treatments with a new PRC1 inhibitor UNC4976, that functions as a chemical surrogate for H3K27me3 and engages chromodomain-containing subunit CBX7. This histone prosthetic inhibition strategy has enabled the discovery that hundreds of chromatin loop and contact domains are repressed by the chromodomain function of PRC1. Repressive chromatin architecture that is dependent specifically upon regulatory function of the PRC1 chromodomain is rapidly de-repressed with chemical inhibition within 4 hours. With Hi-C experiments after rapid drug treatment, and defining drug-sensitive chromatin by ChIP-seq before major transcriptional changes occur, we report canonical PRC1-dependent chromatin architecture in human T lymphocytes. SOURCE: Keji ZhaoLaboratory of Epigenome Biology NHLBI,NIH