Pluto Bioinformatics
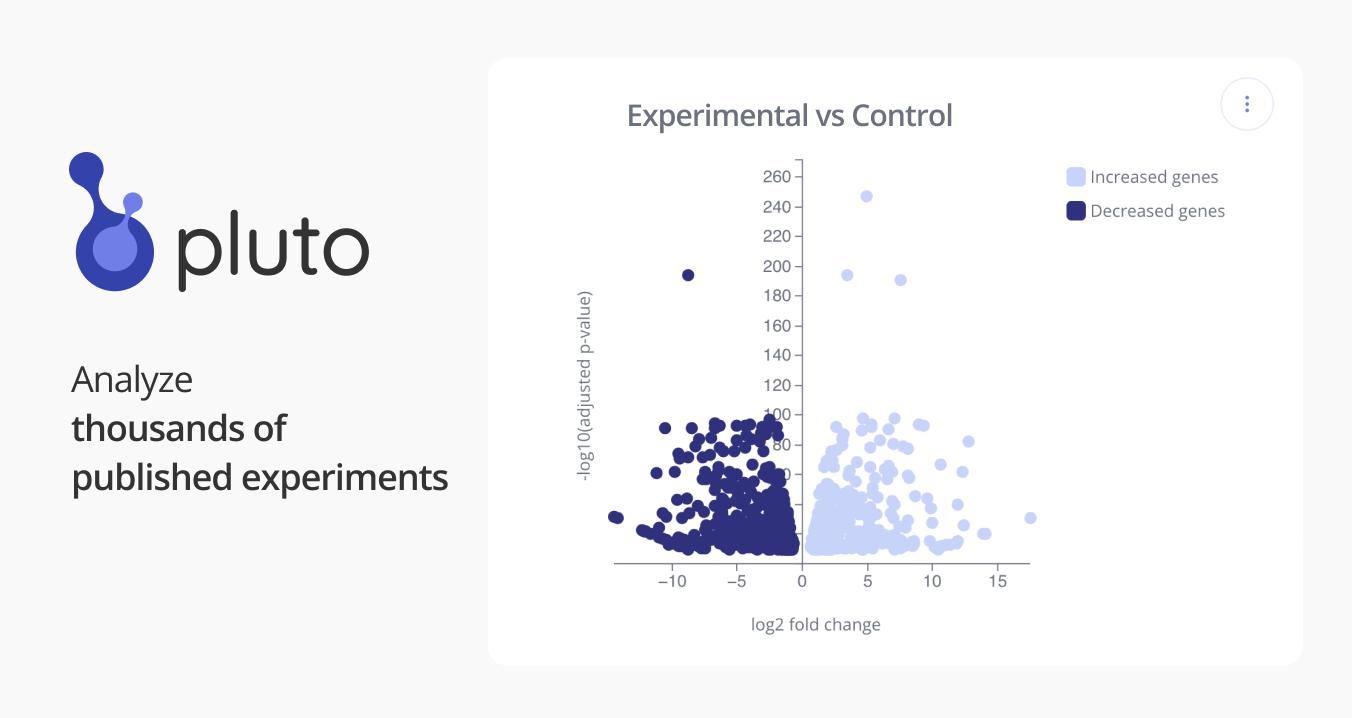
GSE128519: Existence of human-virus chimeric proteins generated during infection
Bulk RNA sequencing
Influenza A virus (IAV) is a threat to mankind because it generates yearly epidemics and poorly predictable sporadic pandemics with catastrophic potential. Influenza has a small RNA genome (~14 Kb) composed of 8 mini-chromosomes (segments). Segments encode both structural proteins and proteins expressed only during infection. Segments are constituted by a 5UTR followed by a coding region and a 3UTR. Transcription of IAV RNA into mRNA depends on host RNA Polymerase II, as the viral polymerase cleaves 5 capped cellular nascent transcripts to be used as primers to initiate mRNA synthesis. We hypothesized that host nascent transcripts bearing AUG could generate upstream ORFs in the viral segments, a phenomenon that would depend on the translatability of the viral 5UTRs. Using orthogonal datasets we report the existence of this mechanism, which generate host-virus chimeric proteins. We show that most segments encode proteins in this manner, expanding the proteome diversity of IAV in infected cells. Host-virus chimeric proteins are conserved across IAV strains, pointing to an evolutionary conservation of function achieved by sampling of the evolutionary space before gene fixation. Thus, we discover a mechanism that generate human-virus chimeric proteins during infection. SOURCE: Max Chang (mchang@ucsd.edu) - University of California, San Diego