Pluto Bioinformatics
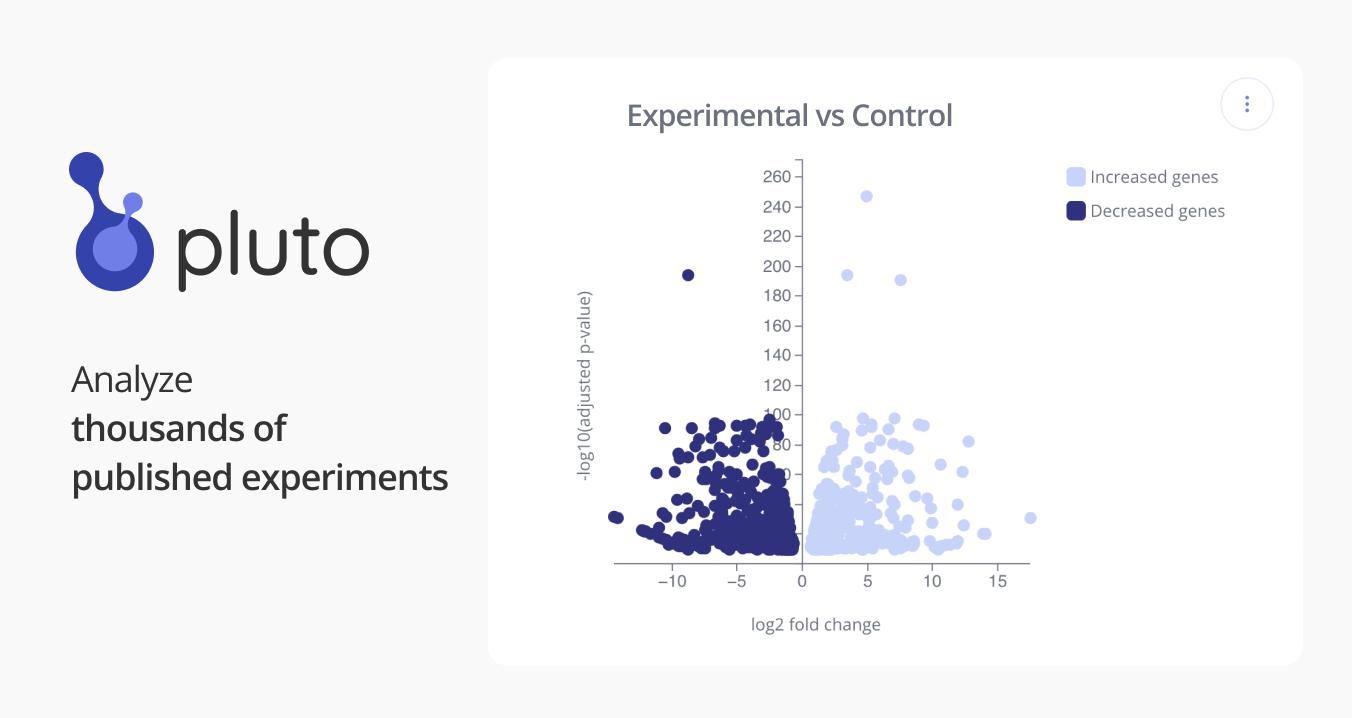
GSE68671: Differential regulation of alternatively polyadenylated isoforms
Bulk RNA sequencing
Alternative Cleavage and Polyadenylation (APA) plays a crucial role in the regulation of gene expression across eukaryotes. Although APA is extensively studied, its regulation within cellular compartments and its physiological impact remains largely enigmatic. Here, we employed a rigorous subcellular fractionation approach to compare APA profiles of cytoplasmic and nuclear RNA fractions from human cell lines. This comparison allowed us to extract APA isoforms that are subjected to differential regulation and provided us with a platform to interrogate the molecular regulatory pathways that shape the APA profiles in the subcellular locations. Subsequently, we depleted these cells of Dicer to compromise the miRNA biogenesis pathway to isolate the differentially regulated APA events that were regulated by miRNA. SOURCE: Jonathan NeveFurger Oxford University