Pluto Bioinformatics
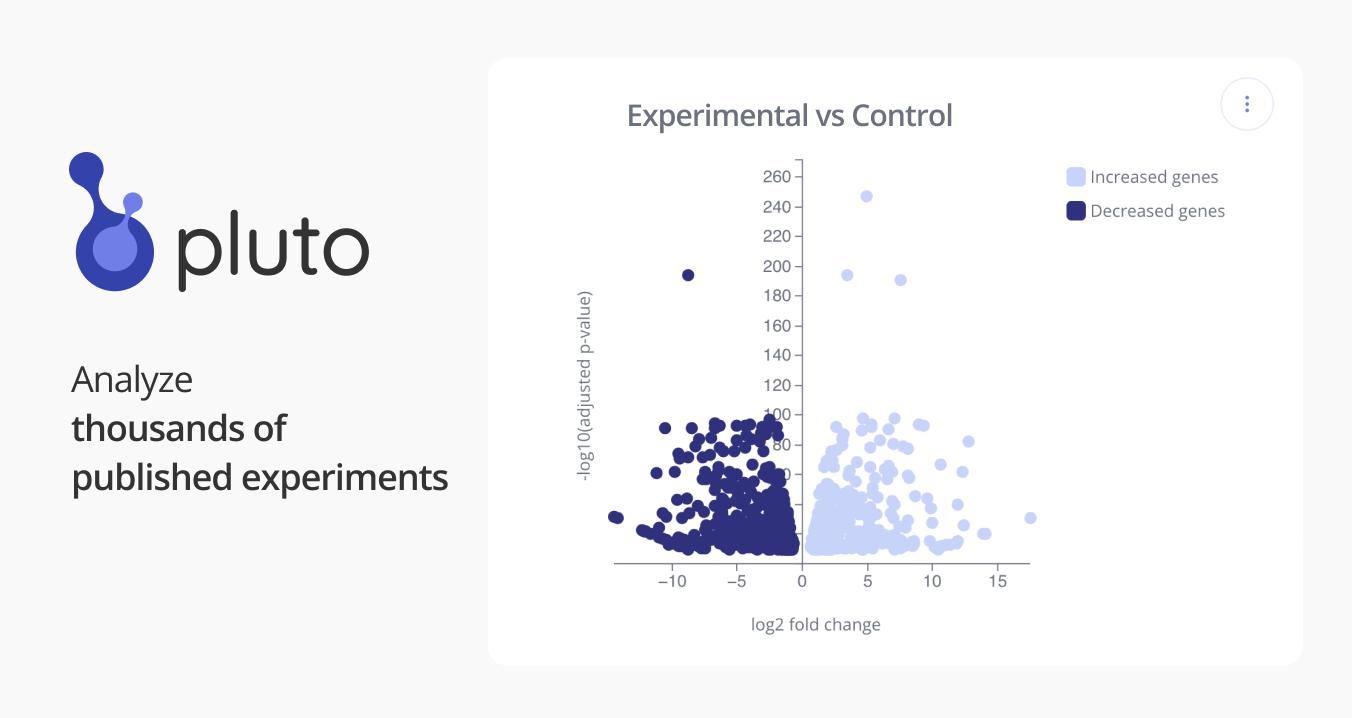
GSE53837: Allelic expression mapping across cell lineages reveal repressor disruption among disease SNPs
Bulk RNA sequencing
Most variants associated to diseases are located in non-coding regions of the genome, and are thought to be regulatory in nature. Cis-regulatory SNPs (cis-rSNPs) impacting transcription can be identified through the mapping of differences in allelic expression (AE). We mapped common cis-rSNPs of protein coding and non-coding genes in 3 distinct cell-types. We show 70% sharing across tissues and similar genetically controlled transcription for protein-coding genes and lincRNAs. Candidate cis-rSNPs altering the expression of 42 non-coding RNA overlap SNPs underlying GWAS associations for 39 diseases. We uncover a new class of cis-rSNPs leading to disruption of footprint-derived de novo motifs, predominantly bind by repressive factors and implicated in disease susceptibility through overlaps with GWAS SNPs. Finally, we provide proof-of-principle for a new approach for genome-wide functional validation of transcription factor SNP interactions. We perturbed NFB action in lymphoblasts and identified 489 cis-regulated transcripts with altered AE after NFkB perturbation. Altogether, we performed a comprehensive analysis of cis-variation in four cell-populations, and provide new tools for the identification of functional variants associated to complex diseases. SOURCE: Tony Kwan (tony.kwan@mcgill.ca) - McGill University