Pluto Bioinformatics
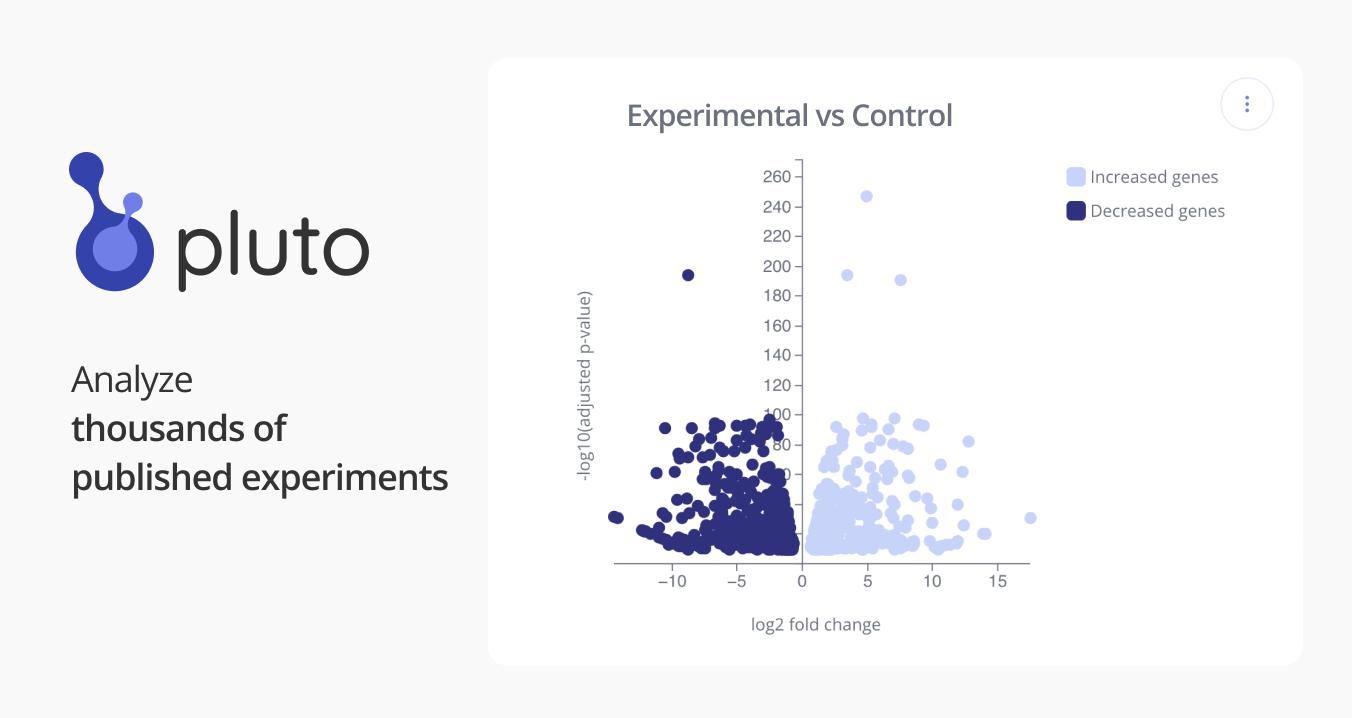
GSE49890: RNA-Seq and expression microarray highlight different aspects of the fetal transcriptome [RNA-Seq]
Bulk RNA sequencing
The second trimester fetal transcriptome can be assessed based on cell-free RNA found within the amniotic fluid supernatant. The objective of this study was to compare the suitability of two technologies for profiling the human fetal transcriptome: RNA-Seq and expression microarray. Comparisons were based on total numbers of gene detected, rank-order gene expression, and functional genomic analysis.; Fewer gene transcripts were observed using RNA-Seq than microarray (4,135 v 8,841). Correlation of total expression within each sample ranged from R=0.43 to R=0.57. On average, there was 59% concordance in gene identity among the top 10% of genes ranked by expression. The RNA-Seq data yielded more significant pathways enrichment within the Physiological Systems Development and Function categories of IPA. Alternative splicing of many well-known genes, including those previously studied in fetal development, such as H19 and IGF2 is detected by RNA-Seq. Also included in this paper is discussion of the technical challenges inherent to working with cell-free fetal RNA and possible solutions. SOURCE: Lillian Zwemer (Lzwemer@tuftsmedicalcenter.org) - Tufts Medical Center