Pluto Bioinformatics
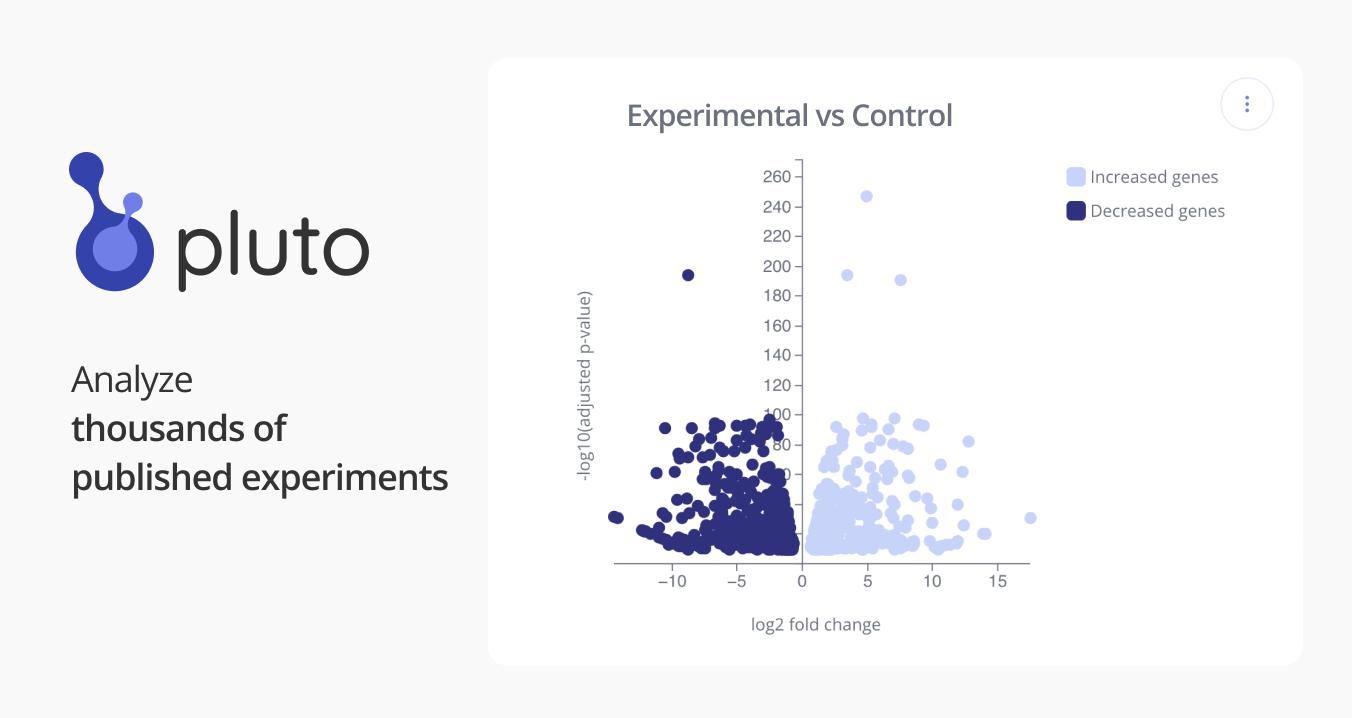
GSE74900: Transcriptome-wide analysis of gene expression regulated by LSD1 during brown adipocyte differentiation [RNA-seq]
Bulk RNA sequencing
By screening a collection of epigenetic compounds, we find that Lysine-Specific Demethylase 1 (LSD1) inhibitors repress brown adipocyte differentiation. RNAi-mediated Lsd1 knockdown shows similar effect, which can be rescued by expression of wild-type, but not catalytically inactive, LSD1. Furthermore, adenoviral Cre-mediated LSD1 deletion in mice leads to inhibition of brown adipogenesis, validating the pivotal role of LSD1 in brown fat development in vivo. LSD1 is a histone H3 demethylase, which selectively removes methyl groups from mono- and di-methylated lysine 4 (H3K4me1 and H3K4me2) under most circumstances, and only from lysine 9 (H3K9me1/2) when bound with androgen receptor (AR) or estrogen receptor (ER). K4 demethylation causes transcription repression, while K9 demethylation may lead to activation of gene transcription. To investigate the target genes of LSD1 during BAT differentiaiton, we performed RNA-seq to profile the gene expression in brown adipocytes treated with DMSO or LSD1 inhibitor 611 (Cpd A) for 6 days, and gene set enrichment analysis (GSEA) was then employed to identify Gene Ontology (GO) terms that were significantly enriched. SOURCE: Wei Qi (vicky.qi@novartis.com) - China Novartis Institutes for BioMedical Research