Pluto Bioinformatics
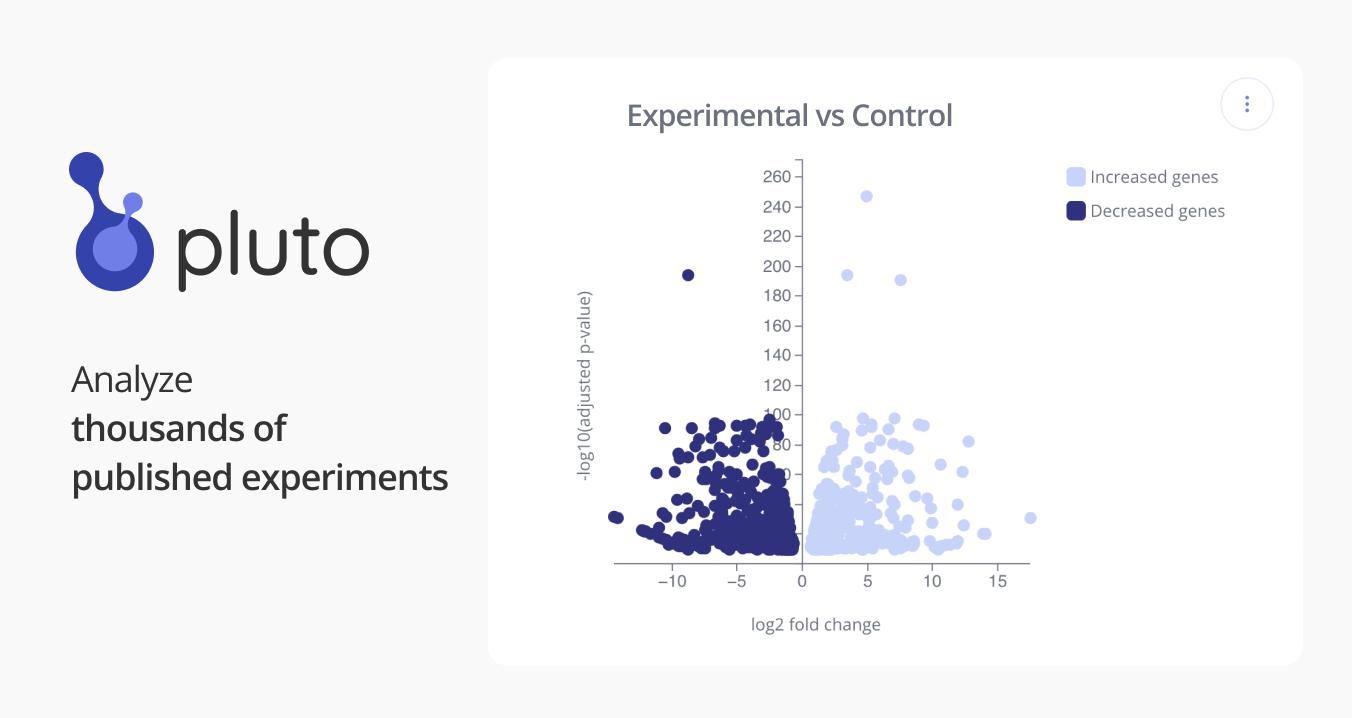
GSE67382: BAF controls genome accessibility
Bulk RNA sequencing
Somatic differentiation requires induction of lineage specific genes to fulfill specialized tissue functions yet the genomic control of this process is incompletely understood. Using the epidermis as a model, we show here that the BAF chromatin remodeling complex is essential to maintain a subset of open chromatin regions, which are strikingly enriched for the DNA binding motif of the stratified epithelial lineage-determining transcription factor p63 (p=1X10-1786). These p63 sites are intrinsically inaccessible. BAF is required to displace nucleosomes ~40 base pairs away from p63 motifs to enable p63 binding, to maintain active histone marks, and to recruit RNA polymerase II to promote gene expression. In this process, p63 was itself also essential for full BAF recruitment to p63 binding sites. Intrinsically, p63 binding sequences favor nucleosome occupancy. Our data therefore suggest a model where a lineage-specific transcription factor cooperates with the BAF complex to establish genome accessibility and engage tissue-specific gene expression. SOURCE: Kun Qu (kqu@stanford.edu) - Paul Khavari Stanford University